When doctors and scientists want to see inside a body, magnetic resonance imaging (MRI) is a powerful tool. MRI can noninvasively capture detailed images of the body’s muscles, organs, and bones. It can monitor blood flow to generate a map of brain activity. And with new sensors developed by bioengineers at MIT, MRI can track the kinds of molecules that make our brains and bodies work.
In the May 13 issue of the journal Nature Biomedical Engineering, a team led by Alan Jasanoff, the Eugene McDermott Professor in the Brain Sciences and Human Behavior at MIT, reports on their new sensors, which can brighten or dim MRI signals in response to specific molecular targets. The probes are designed to amplify the effect that each target molecule has on MRI signal, dramatically improving sensitivity over previous small-molecule sensors. Jasanoff, who is also an associate investigator at the McGovern Institute for Brain Research, says the approach his team used should enable the development of MRI sensors that detect neurotransmitters and other important molecules in the brain.
“We want to be able to measure distinct chemical signals like neurotransmitters, neuropeptides, and metabolites as they fluctuate across the whole brain,” Jasanoff says. “These chemicals are important ingredients in neural computations, and we want to use the types of probes that we developed to detect these signals dynamically.”
Jasanoff explains that researchers have struggled to use MRI to sensitively detect small molecules in the brain because the amount of any given neurochemical is low. Sensors can be designed to change the brightness of an MRI signal in the presence of specific molecules — but it takes a lot of contrast agent to achieve this. If every molecule of contrast agent needs its own target molecule to activate it, low concentrations of the target molecule limit the sensors’ visibility in an MRI scan. “The signal change that you see in the imaging will be very modest,” Jasanoff says. “It won’t let us detect physiological events.”
The Jasanoff team’s new sensors, whose development was led by postdoc Sayani Das and graduate student Jacob Cyert Simon, overcome this problem. To generate a greater signal change in response to target molecules, the researchers designed probes in which a single target molecule impacts not one contrast agent, but many.
To achieve this, Das and Simon packaged an MRI contrast agent inside tiny sacs called liposomal nanoparticles. Each nanoparticle is packed with many molecules of gadolinium, a magnetic material that brightens the MRI signal that arises from hydrogen atoms in water. Inside their protective sacs, gadolinium has no effect on MRI signal, unless water molecules can easily get in and out.
Das and Simon built water channels into the walls of their gadolinium-filled nanoparticles, engineering them so that their opening depends on the presence or absence of a target molecule. When the channels open, more water enters and the gadolinium brightens the local MRI signal, lighting up that spot in a scan.
The researchers call their target-responsive sensors liposomal nanoparticle reporters, or LisNRs (pronounced “listeners”). They designed LisNRs that let water in only in the presence of their target molecule. The water channels in these nanoparticles stay blocked until they encounter their target, which can knock aside a channel-blocking bit of protein.
Once the channel blocker is displaced, water enters and MRI signal brightens. They also made LisNRs that dim the MRI signal in the presence of the molecule they are designed to detect. These have a channel that stays open until the target molecule comes along and blocks it, keeping water out. Jasanoff lab members Vinay Sharma, Samira Abozeid, and Gregory Thiabaud played key roles in understanding and optimizing these interactions, and collaborators in the laboratory of Masayuki Inoue at the University of Tokyo helped the group engineer channels with higher potency.
In experiments led by postdoc Miranda Dawson, Jasanoff’s team used their LisNRs to detect a molecule called biotin in the brains and bodies of living rats, illustrating the probe’s amplifying effects. “We showed that we could detect micromolar-scale levels of biotin with about tenfold greater sensitivity than we would have if we’d used a more conventional, one-to-one type sensing approach,” Jasanoff says. He adds that the team’s modeling suggests that with further development, they may be able to achieve even greater sensitivity gains.
The group showed that the new sensors can be delivered systemically, reaching various organs and spreading throughout the brain. This makes them promising tools for brain-wide imaging, as well as imaging targets in the peripheral nervous system or other tissues.
A next step will be engineering LisNRs that respond to the specific neurochemicals that Jasanoff and his team hope to study. “There are something like 100 neurochemicals in the brain that we’d love to detect, in principle,” he says. They’ll start with dopamine and glutamate — two important and relatively abundant molecules that mediate communications between neurons.
This research, including support for postdoctoral fellows and graduate students involved in the work, was funded, in part, by Lore Harp McGovern, the Yang Tan Collective at MIT, the K. Lisa Yang Brain-Body Center at MIT, the Hock E. Tan and K. Lisa Yang Center for Autism Research at MIT, and the K. Lisa Yang and Hock E. Tan Center for Molecular Therapeutics at MIT.
Place-based pathways to a viable futureLiving Climate Futures Symposium explores climate challenges and solutions at the community level.Aiming to transition away from fossil fuels and avert the worst consequences of climate change, world leaders aspire to achieve net zero global greenhouse gas emissions by 2050 and cap global warming at 1.5 degrees Celsius. But actions to meet such targets and minimize adverse impacts on lives, livelihoods, and infrastructure are not one-size-fits-all; they will require different approaches in different places.
To better understand the patchwork causes and effects of the climate crisis and elements of viable solutions to it, researchers in MIT’s Living Climate Futures (LCF) initiative — 20 MIT faculty and affiliates from across the Institute — collaborate with frontline communities in diverse physical and socioeconomic landscapes around the world.
Funded by the MIT Human Insight Collaborative (MITHIC) and based at the MIT School of Humanities, Arts and Social Sciences (SHASS), LCF is a multi-disciplinary research hub and community of practice; focuses on how climate change impacts people’s everyday lives; and creates knowledge and research collaborations with community organizations.
At MIT on April 23-25 — just after Earth Day — LCF showcased several of these collaborations at its second Living Climate Futures Symposium, which brought together community environmental organizations with MIT researchers and students to explore how climate change challenges and responses to them are playing out in locations from New England to Mongolia.
“Across the next two days, we’ll have conversations about community-based work and scholarly research that’s aimed at understanding the structural causes and social effects of climate change as it’s experienced in people’s everyday lives,” said MIT professor of anthropology and MITHIC faculty co-lead Heather Paxson in remarks at the start of the first full day of the conference. “I’m really excited for this symposium, and for where Living Climate Futures can go from here.”
Resisting environmental harm: Confronting data centers
A session on data centers, energy concerns, and community health in Greene County in Western Pennsylvania highlighted how stakeholders are attempting to proactively avert long-term threats to the environment and public health in and beyond their neighborhoods. Nicholas Hood, senior organizer at the Center for Coalfield Justice (CCJ), which has worked to improve policy and regulations on fossil fuel extraction and use in the region since 1994, described local environmental and health impacts of these activities, including fracking, which has increased water pollution, asthma, and lymphoma. “We have coal mines, these old oil wells, and fracking on top of that, and now we’re going to add data centers,” he said. “So, ask yourself, do you think we want that?”
CCJ community advocate Jason Capello noted that market forces compel data center developers to build as cheaply as possible in places where they believe the population is unlikely to raise concerns about adverse environmental and health impacts. These impacts include pollution from on-site water-based cooling systems, diesel generators and mini-power plants that run on natural gas, and fine particulate matter-linked illnesses such as childhood asthma, heart attacks, stroke, and lung disease. But in a subsequent presentation, Livia Garofalo, a cultural and medical anthropologist on Data and Society’s Trustworthy Infrastructures team in Philadelphia, showed that many communities have pushed back against data center project proposals. “Through protests, canvassing, petitions, and public hearings, communities have been able to resist and even stop data center projects,” she said.
To help communities resist or limit the impact of proposed data center projects, Michael Cork, a postdoc in biostatistics at the Harvard T.H. Chan School of Public Health, described a tool he has developed to estimate emissions, model how pollution would spread, estimate who will be exposed, and assess likely health and economic impacts. To further explore how communities can respond to such projects, MIT associate professor of anthropology Amy Moran-Thomas and Stanford University postdoc Anjuli Jain Figueroa facilitated an educational game conceived by Northeastern University associate professor of sociology and health science Sara Wylie.
The game helped teach participants how often-overlooked community stakeholders can negotiate community benefit agreements (CBAs), or plans that specify project developers’ commitments to address their concerns and provide local improvements such as jobs and affordable housing. Gathered around several tables, symposium participants worked together to identify potential pros, cons, and trade-offs of allowing a data center to be built in a fictitious community. Offering another avenue for community advocacy, Moran-Thomas also moderated a workshop led by public anthropologist Ieva Jusionyte on how to write op-eds that inspire change.
Repairing environmental harm: More than a matter of money
A session on global perspectives and methodologies for potential climate reparations focused on the context for and definition of the term. Veronica Coptis, senior advisor at Taproot Earth, a U.S.-based nongovernmental organization, described her view of climate justice as a movement about reducing not only excessive greenhouse gas emissions, but also changing the systems that have produced them, all while building a world where everyone can live, rest, and thrive in the places they love. “[Taproot Earth’s] mission is building power and cultivating solutions with frontline communities to advance climate justice through Black liberation, Indigenous sovereignty, and democracy,” said Coptis.
Eliane Lakam, global policy and partnerships specialist at Taproot Earth, described a two-decades-long process, sparked by Hurricane Katrina’s devastation of marginalized communities on the U.S. Gulf Coast, that led to a Global Climate Reparations Working Statement at the Global Climate Reparations Governance Assembly of 200 climate leaders in Nairobi, Kenya, in 2024.
Urban agriculture: Reclaiming and revitalizing degraded land
A session on advancing urban agriculture in a changing climate featured a panel of four organizational representatives of various growing spaces in Greater Boston, many of which were formerly vacant lots and garbage dumps that were repurposed as farms and gardens. The panel included Sabrina Pilet-Jones, urban farm manager at Haley House; Cecilia Del Cid, director of food justice and youth programs at GreenRoots; Olivia Golden, urban agriculture educator at UMass Extension; and Matthew Ellison, assistant farm manager at the Urban Farming Institute.
The panelists showed how their efforts to grow food locally in an urban setting are challenging past and ongoing environmental inequality in myriad ways. These include preserving and expanding green spaces, increasing access to fresh produce, empowering their communities to become actively engaged in how their food is grown, building community connection and pride, and inspiring young people to grow food in their neighborhoods. They framed their organizations’ youth education programs as gateways for enabling the transfer of knowledge from elders to young people, promoting a strong work ethic and healthy lifestyles, and identifying pathways to livelihoods that address food access and sustainability. To provide participants with an opportunity to learn about urban agriculture and do some volunteer farm labor, the symposium offered a field trip to The Food Project in Roxbury.
Rural and urban adaptation: Responding to a changing climate
A session on climate change as a place-based phenomenon explored how communities are responding to a changing climate on Mongolian grasslands, in the greater Southwestern United States, and along the Boston Harbor.
Munkh-Erdene Gantulga, a PhD candidate in geography at the School of Geography and the Environment at the University of Oxford, described his studies at the National University of Mongolia on how pastoralists at two field sites are protecting their livelihoods as more-frequent severe weather events increase livestock mortality and pasture degradation. Perceiving climate change as a lack of rainfall, hotter temperatures, and inadequate grass growth, herders at the two sites are either migrating to greener pastures or applying three strategies: not milking their animals so as to boost survival of mothers and their offspring; selling off parts of their herds; or specializing in more climate-resilient animals, such as camels. A separate screening of the film “If Only I Could Hibernate” dramatized the environmental and economic obstacles faced by youth in Mongolia.
Breanna Lameman, an Indigenous data sovereignty doctoral scholar and graduate research associate at the University of Arizona, and Nekai Eversole, wildlife biologist and program lead with Climate Change Program - Navajo Nation Department of Fish and Wildlife, described how traditional Diné ecological knowledge and innovative technologies are helping Navajo Nation communities to adapt to hotter temperatures, long droughts, and harsher soil conditions. Lameman cited Diné concepts of restoring balance and maintaining kinship with the natural world as essential to the local response. “This reminds us that the plants, animals, water, and soils are relatives, not resources, and that we all need to work together,” she said. “Watching the stars, observing the winds, the plant cycles, and animal behaviors, really helps us predict seasonal shifts better than any app out there.” Eversole noted that this mindset is combined with innovative technologies ranging from hydroponics to wetland restoration structures. A separate screening of the film “Climate Voices” and Q&A with director Leslie Jonas, MLK Jr. Visiting Scholar and Elder Eel Clan member of the Mashpee Wampanoag Tribe, explored perspectives from Native experts and climate scientists working on the front lines.
Elisa Guerrero, community engagement manager at the Stone Living Lab and Sustainable Solutions Lab at the University of Massachusetts Boston, highlighted two examples of adaptation measures to protect vulnerable Boston Harbor infrastructure from sea-level rise, coastal storms, and storm surges: testing seawalls designed to mimic natural habitat for how well they slow down wave action and preserve marine biodiversity, and monitoring salt marshes to better understand the factors that degrade and promote their health. A separate Stone Living Lab tour enabled symposium participants to visit a living seawall, nature-based flood protection infrastructures, and a community-based flood sensor project as Boston tries to address rising sea levels.
Training the next generation in community-oriented research
In addition to highlighting LCF’s role as a research hub linking MIT researchers and students with community organizations in the United States and around the world, the symposium also sought to draw attention to efforts to train the next generation in this approach. The Saturday session “Experiential Learning, ‘Anthro-Engineering,’ and Learning to Do Community-Oriented Research” showcased some of the interdisciplinary classes that LCF supports. MIT students who participated in these classes engaged in activities ranging from building chicken coops with a Boston farming collective while learning about urban agriculture to exploring how to decarbonize the steel industry in Pittsburgh and Southeast Chicago while creating well-paying green jobs to spending time in Ulaanbaatar’s ger districts (informal residential areas) while working with Mongolian collaborators on non-coal methods for heating homes.
Student panelists shared highlights from their learning experiences through presentations, activities, artwork, and written accounts from their travel notebooks.
“People have always been part of why I chose to study engineering,” said nuclear engineering PhD student Alina Jugan. “But learning how to integrate a human perspective, and one that accounts for multitudes of realities, is essential. The first step in making a solution is learning what the real problem is and how people experience it. This is what ‘Anthro-Engineering’ teaches us.”
Panel and symposium co-organizer Laura Frye-Levine, a research scientist at the MIT Anthropology Section and affiliate of the MIT Center for Sustainability Science and Strategy, concurred. “In building relationships in place-based contexts, the students on this panel demonstrate the value of engaging with social and cultural expertise in addressing climate change,” she said. “These projects are fantastic examples of collaborations that hold promise for MIT’s approach to developing climate solutions.”
Lessons in resilience from frontline community groups
In a session entitled “Xa xah Xechnging: A Sacred Obligation in a Time of Climate Chaos,” panelists from Se’Si’Le and Children of the Setting Sun Productions — two Indigenous-led environmental organizations from the U.S Pacific Northwest that have collaborated with LCF on experiential learning activities — described how they draw upon cultural, spiritual, scientific, legal, and other resources in their efforts to heal and restore the planet amid political and corporate opposition. At the core of their work is a perspective in which everything has a spirit, and is thus worthy of love, honor, respect, dignity, pride, and compassion.
Sundance chief Rueben George, a board member of Se’Si’Le, recounted how this perspective energized the campaign he led against the development of the Trans Mountain Pipeline, a fossil fuel megaproject on Tsleil-Waututh Nation territories in British Columbia. “We just shared facts about what it is, and we led with our culture,” said George, who is also chair of Salish Elements, an Indigenous-run company that produces green hydrogen. “That’s the biggest, most important thing, is we always led with our culture.”
At an earlier session, representatives of organizations that participated in the 2022 Living Climate Futures symposium, ranging from GreenRoots to Se’Si’Le, said that they draw strength from the wisdom of ancestors, a growth mindset, and communal bonds among people who seek a better future for the places they call home. Kurt Russo, co-executive director of Se’Si’Le, noted: “I come back to the indomitability of the human spirit.”
Additional photos can be viewed here.
Ten from MIT accept 2026 Fulbright awardsThis year, over half of MIT’s Fulbright applicants won awards. The current students and alumni will embark on research projects abroad in 2026-27.Ten MIT affiliates — including undergraduates, graduate students, and alumni — have accepted Fulbright grants to conduct research in countries across the world. Five other students declined their awards to pursue other opportunities, and another student is still deciding. In total, 16 of MIT’s 30 Fulbright applicants won awards this year.
Funded by the U.S. Department of State with annual appropriations from Congress, the Fulbright U.S. Student Program offers year-long opportunities for American-citizen students and recent alumni to conduct independent research, pursue graduate studies, or teach English in over 140 countries. This past February, MIT was recognized by the Fulbright Program as the nation’s No. 1 “Top Producing Institution” among special focus STEM universities.
MIT students and alumni interested in applying to the Fulbright U.S. Student Program should contact Julia Mongo, Fulbright program advisor, in the Distinguished Fellowships office in Career Advising and Professional Development.
Jessica Chomik-Morales SM ’25 earned her master’s in science writing at MIT, where she previously spent three years as a post-bac cognitive neuroscience researcher in the labs of professors Nancy Kanwisher and Laura Schulz. For her Fulbright in Spain, she will research the science of science communication at Universitat Pompeu Fabra’s Center for Brain and Cognition in Barcelona. Her project will investigate how narrative features in science writing interact with reader characteristics to shape comprehension, trust, and engagement. Chomik-Morales is the creator, host, and producer of “Mi Última Neurona,” an MIT-sponsored Spanish-language neuroscience podcast that has featured more than 60 scientists from Latin America, the United States, and Spain. She is currently producing “Lab Notes on Love,” an audio miniseries for Scientific American. She is committed to making science communication more inclusive, empirically grounded, and emotionally resonant.
Stella Gassman will graduate this month with a BS in biological engineering and a concentration in women’s and gender studies. For her Fulbright year, she will conduct microbiology research at the University of Copenhagen in Denmark. At MIT, Gassman researched the vaginal microbiome and mucosal membranes, with a particular focus on bacterial vaginosis. Important moments of her research journey included time at an MGH gynecology clinic and at the FRESH clinical trial site in South Africa, where she gained firsthand perspectives on the human context behind her laboratory samples. Gassman also interned at Pfizer Oncology, developing an in vivo tumor model to test preclinical compounds. She volunteered in the MGH Emergency Department and served on the Biological Engineering Undergraduate Board. After Fulbright, she hopes to attend medical school to bridge scientific discovery and human impact.
Chen Li SM ’25 graduated from MIT with a master’s in system design and management. She has developed generative artificial intelligence tools for patient engagement at Novo Nordisk in Copenhagen through MISTI Denmark and applied AI to help prevent gait freezing in Parkinson’s patients through the MIT–Mexico program. As a research assistant in the MIT Global Teamwork Lab, her thesis used large language models and statistical methods to build a 3D urban design platform to study teamwork behavior. She also served as a teaching assistant for data mining courses at MIT Sloan School of Management and the MicroMasters program. As a Fulbright Iceland-NSF Arctic Research Award recipient, Chen will explore how AI and systems thinking can be applied to support health and well-being in Arctic communities. She plans to pursue a PhD in information and systems science following her Fulbright experience.
Liam Moser will graduate this week with a PhD in geophysics from the Department of Earth, Atmospheric and Planetary Sciences’ MIT-WHOI Joint Program. His research has focused on understanding the structure and dynamics of subduction zones, where one tectonic plate dives beneath another, generating the Earth’s largest earthquakes and creating volcanic arcs. During his PhD, Moser helped found the annual MIT-WHOI Geophysics Retreat, promoting interconnectedness between MIT and the Woods Hole Oceanographic Institution (WHOI). He also taught incoming graduate students in the MIT-WHOI Summer Math Review for five years, organizing the review for the final two years of his PhD. Moser was awarded a Fulbright Iceland-National Science Foundation Arctic Research Award for a postdoctoral fellowship at Reykjavík University, where he will use earthquake recordings to study the structure and dynamics of the Hengill volcano and geothermal area.
Lilia Ould-Hammou is a senior majoring in mechanical engineering with a concentration in controls, robotics, and instrumentation. As a recipient of the Fulbright U.S.-Korea Presidential STEM Initiative Award, she will conduct research at Seoul National University’s Wearable Robotics Laboratory. Her work will involve advancing adaptive exosuit control for balance recovery. She plans to improve her language skills while exploring Korea’s history and culture. At MIT, Ould-Hammou has worked in the d’Arbeloff Robotics Lab on soft modular robotic straps, served as a tutor in the MIT Women’s Technology Program, and competed as a thrower on the MIT track and field team. After her Fulbright fellowship, she will pursue a master’s degree in robotics at Johns Hopkins University.
Bryan Sperry ’23 graduated from MIT with dual bachelor’s degrees in physics and mechanical engineering, focusing on renewable energy systems. Since graduating, he has worked at VEIR as a systems integration engineer, designing superconducting power transmission lines. As a Fulbright Brazil grantee, he will study pathways to improve climate resilience and energy equity in urban power grids alongside the Cenergia Lab at the Federal University of Rio de Janeiro. After Fulbright, he plans to enroll at Columbia University to complete a master’s in urban planning to continue working on urban disaster preparedness.
Sophie Thompson is a senior majoring in chemical engineering. For her Fulbright research in Sweden, she will test the performance of recycled carbon fiber composites at the Swedish School of Textiles in Boras. Thompson has researched natural fiber-reinforced composites for prosthetic socket use in low-resource environments with the Herr Lab in the MIT Media Lab, worked on immunoengineering technology at the Massachusetts General Hospital, and interned at the textile recycling startup MacroCycle Technologies. She also completed a summer research internship at the Weizmann Institute in Israel through MISTI. She serves as captain on the MIT lightweight women’s rowing team, and has held leadership roles with the MIT chapter of the American Institute of Chemical Engineers, TEDxMIT, and MIT Hillel. After Fulbright, Thompson will pursue a PhD in molecular engineering at the University of Chicago.
Claire Underwood is a senior studying chemical-biological engineering. As a recipient of a Fulbright Portugal award, she will conduct research at the University of Minho in Guimaraes, studying high-throughput fabrication techniques for cell-embedded microtissues with applications in drug discovery. At MIT, Underwood worked in the Hammond and Olsen labs exploring interactions between biology and polymeric systems. For the past two years, she has focused on lipid nanoparticle drug delivery for cancer treatment, and is excited to continue investigating biomaterials and biomimetic systems. She was also a member of the varsity volleyball team and active in her sorority Alpha Phi, Cru, and Athletes in Action. After Fulbright, she will pursue a PhD in chemical engineering at the University of Texas at Austin.
Sophie Vulpe is a senior majoring in physics and mathematics. Her Fulbright will take her to the Extreme Light Infrastructure-Nuclear Physics (ELI-NP) institute in Măgurele, Romania, where she will develop advanced data-processing algorithms for a new monoenergetic gamma ray spectrometer. She looks forward to strengthening her computational and experimental skills and connecting with her Romanian heritage. At MIT, Vulpe worked with Professor Mikhail Ivanov on characterizing black hole quasi-normal modes using tools from the mathematical field of representation theory. Passionate about expanding access to physics through education and outreach, she was co-president of the Undergraduate Women in Physics group, a mentor in the physics mentorship program, and a teaching assistant in the Experimental Study Group. She was also a member of Dancetroupe and the Musical Theater Guild. After Fulbright, Vulpe plans to pursue a PhD in physics.
Josephine Wang will graduate this month with a BS in computer science. For her Fulbright grant to Switzerland, she will conduct research at EPFL in Lausanne with the NeuroAI Lab. Her work will explore whether brain-inspired language models can develop functionally specialized clusters analogous to cortical organization, and how targeted disruptions to those clusters affect language-related behavior. At MIT, Wang’s research has focused on computational models of cognition, movement, and human behavior. She has most recently worked in the Seethapathi Motor Control Group, where she developed a computer vision pipeline for world-grounded pose estimation in children and examined how computational models can support pediatric gait analysis. Outside of research, Wang enjoys traveling, trying new cuisines, and learning French.
MIT students study plasma physics beneath Alaska’s auroraStudent-led expeditions use distributed instruments to observe auroral structures and probe space plasma in real-world conditions.For many graduate students, waking up at noon after a 4 a.m. bedtime is a sign of a night well spent. For a group of MIT students, it was simply the start of their workday — timed not to the sun, but to the aurora.
Their goal was simple: to study plasma phenomena using the aurora borealis as a natural laboratory. The process, less so; working largely in darkness in Fairbanks, Alaska, the students conducted experiments in temperatures that dipped as low as -25 degrees Fahrenheit, using red headlamps for visibility. The sun set before 3 p.m., and even at its warmest, temperatures barely reached 20 F.
The aurora provides a rare opportunity to observe plasma behavior directly, as charged particles that interact with Earth’s magnetic field produce visible, large-scale structures in the night sky. As Fairbanks is situated beneath a region of especially frequent auroral activity, it is one of the most reliable places in the world to observe these phenomena, though the conditions come with real constraints.
For one thing, the extreme cold directly impacted the instrumentation. “Our laptops went from full battery to nearly empty in 10 minutes because of the cold,” says Leonardo Corsaro, a PhD student in physics at the Plasma Science and Fusion Center (PSFC) at MIT. “We were trying to transfer data as fast as possible before everything shut down; it was a race against time!”
The challenges extended beyond the cold itself. “The cold can be managed,” says Leon Nichols, a PhD student in physics at PSFC. “With good planning, you can stay comfy in -20 F. The real difficulty was movement when deploying cameras far away from the roads. Walking through thick snow can burn up to 900 calories in an hour. We used cross-country skis to access some of the more remote terrain that would have taken hours to reach otherwise.”
But the conditions were more than worth it: During their time in Alaska, the group witnessed the strongest solar storm in the past two decades, bringing the aurora to life in ways few will ever experience. “It felt like we were the only ones there,” Sydney Menne, a PhD student in nuclear science and engineering, recounts, “removed from the Earth and just entirely surrounded by the aurora, fully immersed in it.”
The team was granted access to observation facilities at Poker Flat Research Range through the University of Alaska Fairbanks Geophysical Institute. Over the course of the trip, students deployed multiple all-sky camera systems across distances of up to 100 miles, enabling simultaneous observations of auroral structures from different locations. These cameras, which capture 360-degree images of the night sky, were paired with magnetometers to correlate visual auroral features with changes in Earth’s magnetic field.
By combining spatially distributed imaging with magnetic field measurements, the team aimed to capture how auroral structures change across space, with the long-term goal of supporting three-dimensional reconstructions of the aurora. This year’s campaign also expanded the measurements beyond imaging, using muon detectors to explore possible correlations between visual auroral activity, magnetic field changes, and particle detections, offering a potential window into how high-energy particles in the upper atmosphere relate to visible auroral activity.
Despite decades of study, many aspects of the aurora remain poorly understood, and each observation offers an opportunity to better characterize the behavior of plasma in near-Earth space. The team also observed a pulsating aurora, a relatively rare phenomenon in which strips of light stretching across the sky blink on and off multiple times per second. By combining instruments not traditionally applied to these problems and deploying low-cost systems at scale, the team is exploring new approaches to studying these phenomena. Insights from these observations can help improve our understanding of space weather, including how solar activity affects satellites, communications systems, and power infrastructure on Earth.
For some participants, the experience reshaped how they think about plasma physics itself. Corsaro explains, “In my research, it is easy to associate these phenomena with colorful plots and simulations, losing touch with the physical process. Seeing structures in the aurora, electric currents and flows forming and shifting overhead, brought a sense of reality to those concepts, and served as a reminder that real plasmas are far less neat and intuitive than theory suggests.”
The experience is part of a broader effort. This group of students represented the third iteration of the Geophysical Plasma Observation Expedition (GPOE), a project involving MIT students from the Plasma Science and Fusion Center, along with collaborating departments, that sends a cohort to Fairbanks, Alaska, each year. Faculty members now provide support for the expedition, while continuity is maintained through its student-driven structure, with each cohort including a mix of returning and new participants. The expedition is organized and led entirely by students and operates on an intensive, compressed timeline. Students are responsible not only for data collection, but also for instrument design, site selection, logistics, and post-processing, completing a full research cycle within a matter of months.
This year’s cohort included graduate students Leonardo Corsaro and Leon Nichols of PSFC; Sydney Menne of the Department of Nuclear Science and Engineering; and Noah Wolfe and Oleksandra “Sasha” Lukina of the Laser Interferometer Gravitational-Wave Observatory (LIGO) Laboratory and the MIT Kavli Institute for Astrophysics and Space Research. The group was accompanied by Professor Matthew Evans, professor of physics at MIT, who is affiliated with the LIGO Laboratory and the Kavli Institute.
“This is an opportunity to go from concept to data analysis in just a few months,” says John Ball, a PhD student in nuclear science and engineering at PSFC. “That kind of compressed scientific cycle is rare, especially in our field.”
The program itself has relatively recent and somewhat unusual origins. It began in 2023, when graduate student Shon Mackie, frustrated by the lack of hands-on plasma diagnostic opportunities, noticed the solar cycle was approaching its peak and saw an opportunity to study plasma phenomena more directly. He drafted a short proposal to PSFC leadership, and the response from then-Director Dennis Whyte was two lines: “Sounds cool, literally! PSFC will fund this.”
Since its launch in 2023, GPOE has evolved from a single-camera effort into a multi-instrument, multi-site campaign with growing participation, with each cohort building on the work of previous years by refining instrumentation, expanding observational coverage, and improving data collection strategies.
This hands-on, student-driven approach has also created opportunities to extend the experience beyond MIT. In 2024, the program expanded to include a new outreach collaboration with the MIT Museum and the MIT Nord Anglia Collaboration, bringing approximately 65 high school students from around 20 schools worldwide to MIT to help design and build components of the all-sky camera systems used in the field. Working within a set of technical constraints, students developed and tested designs, ultimately producing 13 cameras that were deployed during the Alaska expedition.
The program has also begun to produce results beyond the expedition itself. Students have presented their work at major conferences, including the American Geophysical Union, and published findings in peer-reviewed journals such as Earth and Space Science. The group’s low-cost all-sky camera and magnetometer design is now being adopted by other research teams and community science initiatives, extending its impact beyond MIT.
Beyond its scientific goals, participants emphasized the broader impact of the experience.
“Standing outside at midnight in Alaska, staring up at sheets of glowing plasma stretching thousands of kilometers across the sky, really brings home just how small and delicate our own place in the universe is,” says Ball.
As the program continues to grow, students hope to expand both its technical capabilities and its reach, including more permanent instrumentation and expanding outreach partnerships. For many involved, the expedition represents not just a research opportunity, but a reminder of the scale and immediacy of the phenomena they study.
“Science is an adventure,” Corsaro says. “This kind of work reminds you why you became a scientist in the first place.”
Learning to teach, learning to discoverMIT senior Nik Sandu bridges scientific research with a strong commitment to teaching and community.Nik Sandu points to a graph on the whiteboard in a seventh-grade science class. “According to the graph, what is the energy of the ball?” she asks, gently waving a hand to settle the room’s twitchy energy. “Voices are off.”
A student raises his hand, noticing an askew Y-axis. “This one doesn’t go through zero. A ball with no mass shouldn’t have any kinetic energy.” Sandu glances at the board, then back to the class. “Yes, that’s true — that was a little oversight on my part. No mass with no energy is a really good observation.”
The moment captures the MIT senior’s teaching style — reflective, open, and willing to learn alongside her students.
“The No. 1 most important thing in a teacher is being able to evaluate your own teaching, not take it personally, and fix things up as you go,” says Liza Huntington, Sandu’s mentor at the Community Charter School of Cambridge (CCSC). “Nik has that.”
That same openness and on-the-fly adaptability extend beyond the classroom. A materials science major working toward her teaching license through MIT’s Scheller Teacher Education Program (STEP), Sandu has taught high school in Norway, researched magnetoelectric materials and soil stabilization, worked in a Finnish mine extracting ferrochrome, and held leadership positions centered on community building. Each experience reflects a lifelong learner making an impact across roles and settings.
“You can see that in how she tackles different roles, whether it’s research or teaching or service,” says Professor Jeffrey Grossman, her advisor in the Department of Materials Science and Engineering (DMSE). “She’s multidimensional in all the right ways.”
Roots across borders
Born and raised in Chicago to Romanian immigrants, Sandu credits her parents for her commitment to education. They left Romania, where access to higher education was restrictive, so her mother could attend college in the United States.
“She got a bachelor’s in English, which wasn’t her first language,” Sandu says. Her mother later went into the workforce, but still aspires to earn a master’s degree. “Hearing her say, ‘I want to go back to school’ made me think: She’s successful. She came here to America and got her education — and that motivation is a big part of why I’ve come to love education, both as a student and a teacher.”
With limited financial resources, Sandu’s parents strongly supported her intellectual pursuits. Music was a particular focus, a way to stay connected to Romanian culture. “So I know a lot of Romanian songs. When I go home, there’s Romanian music playing.”
By age 6, they encouraged her to take up guitar. She later performed in a youth folk band, the Young Stracke All-Stars, named for Chicago musician Win Stracke. The group played fundraisers and festivals, including an event at Wrigley Field, alongside folk music greats such as Arlo Guthrie and “First Lady of Children’s Music” Ella Jenkins. They also presented Jenkins with a lifetime achievement award and played a tribute concert for Pete Seeger.
Sandu laughs at her younger self’s perspective. “When you’re a kid, you’re like, ‘Oh, this is a cool musician.’ You don’t realize, ‘Oh my God, I met this person.’”
The road to MIT
Sandu became fascinated by STEM subjects at a selective-enrollment public high school in Chicago, taking advantage of every opportunity available, including advanced placement classes, college-credit courses, science fairs, and a class at a local community college.
But many experiences felt out of reach — elite courses and programs she was accepted into, but couldn’t afford. “My family wasn’t really in a position to drop $2,000 on a course or science equipment.”
A school like MIT seemed like a stretch.
She applied anyway on a whim and had the surprise of a lifetime when she saw her acceptance on her phone while in the car with her mom and sister — “I see the confetti and I’m like, ‘Oh my God, I got into MIT,’” she recalls.
She attributes her acceptance to breadth of experience and persistence, but especially to mentors — “teachers who knew how to push me, who knew how to support me, and wanted to see me grow as a student,” she says.
Their influence ultimately led her to the STEP program, which prepares students to become K-12 educators: “I was like, ‘I’m doing this for sure.’”
At CCSC, a tuition-free school that serves many sixth- to 12th-grade students from disadvantaged families, Sandu feels she has come full circle in the classroom.
“Seeing their opportunities and seeing them pushing themselves, and me wanting to be that person who also pushes them to be better students — that has just really affected my worldview,” she says.
A scientific lens
For Sandu, teaching is a way to share her love of science, from geology and the environment — as a first-year student, she wanted to be an Earth, atmospheric, and planetary sciences (EAPS) major — to materials science, while also recognizing how closely the fields connect.
After taking a few EAPS classes, “I was more interested in why these rocks behave the way they do on a chemical or a molecular level.”
That curiosity carried into her 2025 summer internship at Outokumpu Stainless, where she worked in a company-owned mine extracting ferrochrome, a chromium-iron stainless steel alloy. Her coursework helped her understand how the unusually deep, vertical mine formed over millions of years, and how minerals form.
“It was really cool that I had the background to understand that,” Sandu says.
Her research through MIT’s Undergraduate Research Opportunities Program (UROP) followed a similar path — starting with an EAPS study investigating how roots stabilize the soil. She helped choose materials for a 3D-printed root network model and developed a data-collecting procedure using a high-speed camera.
She applied that same problem-solving approach in later materials science research, studying magnetic waves called magnons in a garnet material, tracking how boundaries known as domain walls moved and could be steered in different directions — work with applications in energy-efficient electronics.
Miela Gross PhD ’25, her supervisor on the project, says Sandu’s methodical approach to research was an enormous help. She organized massive datasets, troubleshot equipment issues, and even mastered pulsed laser deposition — a technique for growing new thin films.
What stood out — beyond her research skills — was her facility with communication. Gross described her as clear, proactive, and collaborative; Sandu would raise issues as soon as they came up, rather than waiting to report them later, and she was deliberate about setting and negotiating deadlines depending on her classwork and availability.
“She was really good at asking the right questions: ‘OK, these edge cases that might come up, what should we do there?’” says Gross, now a postdoc at Helmholtz-Zentrum Berlin, a national lab in Germany. “Nik’s passion for science is clear, but her communication is what drives the impact of her work.”
Sense of community
For Sandu, education and community are inseparable. In addition to her research and teaching at CCSC, she has served as a teaching assistant in MIT physics classes, been social chair for DMSE’s Society for Undergraduate Materials Scientists, and helped coordinate the weeklong First-Year Pre-Orientation Program (FPOP) for incoming students, which included a scavenger hunt and an “Among Us”-themed murder mystery, complete with inflatable costumes.
“I prioritize community in all aspects of my life — social, academic, personal,” she says. “For FPOP, we wanted incoming students to know that there’s a community for them at DMSE.”
Gross saw Sandu’s generous spirit extending beyond MIT, noting how she tutored a family friend in high school physics. “She’s just very giving,” Gross says.
Rhea Vedro, lecturer at DMSE and metals artist-in-residence, taught Sandu in class 3.095 (Introduction to Metalsmithing), where students learn to solder and form metal objects. In addition to Sandu’s incredible drive — “if a minimum of five rivets was required, Nik would do 20” — Vedro highlighted a strong commitment to peers, pointing to leadership in the classroom.
“Nik would always do the very best, and that would set a tone for the rest of the group — to rise to that standard,” Vedro says.
Paths forward
Now in her final semester at MIT, Sandu is preparing to pursue a PhD at Dartmouth College, where she will combine materials science and geology to study cobalt minerals critical to lithium-ion batteries. The research builds on her earlier work on magnetism and mineral structure.
At the same time, she is completing the requirements for her Massachusetts and Illinois teaching licenses. It has been a difficult balance between the pull of research and teaching.
“I’m not quite done learning about the world. I’m not ready to leave the academic environment where I’m expected to keep learning,” she says. But teaching remains a long-term goal, especially in Chicago.
“I want to give back to that community that helped me grow, helped me become who I am today,” Sandu says.
Her advisor, Grossman, says Sandu stands out for how she thinks about her experiences and what she learns from them.
“Just hearing her reflect on her experiences — it’s very mature. And that’s a really important part of the experience and the growth that undergraduates do here,” Grossman says. “It’s taking a step back, thinking about what you’ve done, understanding it more deeply, and learning about yourself in the process. Those are the skills of lifelong learners, and she’s already showing them in spades.”
The rules neurons follow to make sense of what we seeBrain cells take in many signals through thousands of circuit connections. A new study discerns the rules that turn inputs into a functional arrangement for neurons that process vision.Even in the primary visual cortex, a brain region named for its specialized role in processing basic features of what the eyes see, not every neuron ends up answering the call to process properties of visual input. Maybe that’s because each neuron receives a wide variety of inputs via thousands of circuit connections, or “synapses,” and has to opt to respond to the visual information versus something else. In a new study in mice, neuroscientists at The Picower Institute for Learning and Memory at MIT reveal how neurons that perform visual processing bring order to this input to get the job done.
Neuroscientists are keenly interested in what inputs, from among so many choices, will compel neurons to participate in the brain’s computations and functions, says senior author Mriganka Sur, Newton Professor of Neuroscience in the Picower Institute and MIT’s Department of Brain and Cognitive Sciences. Neurons ultimately participate in brain circuits by “firing” an electrical action potential.
“The configuration of inputs, the kind of organization, the assembly of neurons that modulate each other to generate an action potential is the essence of how brain circuits process information,” Sur says. “These (visual cortex) cells are a microcosm of this very profound and big picture of neuroscience.”
In the open-access study in iScience, led by postdoc Kyle Jenks, the research team achieved their findings by meticulously imaging how not only neurons’ cell bodies, but also their individual synapses, formed on protrusions known as dendritic spines, responded as mice viewed moving images. They did this imaging for not only visually responsive neurons, but also for unresponsive neurons that nevertheless have visually responsive spines. That allowed them to analyze many key properties that might influence where a particular synapse forms, and how it influences responses at the cell body.
“This pulls together a lot of things that have been looked at in isolation and looks at them in one collective paper,” Jenks says. “We can compare how the neuron and the spines on that neuron respond to the same stimuli, and we can do this for both visually responsive and unresponsive neurons.”
In visual cortex layer 2/3, Jenks and the team genetically engineered neurons such that their individual dendritic spines would glow when surges of calcium indicated increased activity by the synapses on the spines. The scientists did the same for the cell body, or “soma,” to keep track of how the cell responded and even signaled its overall responses back out to the synapses. This way, as the mice watched black and white gratings at varying angles drift by their eyes in different directions, the scientists could keep track of each spine’s and each cell’s overall response to that patterned visual input.
In all, they tracked 11 neurons that responded to the visual input and 11 others that seemingly ignored it. That enabled them to find several rules:
Distance from the soma matters: On cells that responded to visual input, the responses of individual spines were much more likely to correlate with the activity of the soma the closer the spine was to the soma. In the same vein, the soma’s signal back out to spines, which is believed to influence the spines’ alignment with the soma’s preferences, was more likely to be detectable closer to the soma than farther away.
Local clustering: On neurons that responded to visual input, spines formed distinct little enclaves of correlated responses with each other. Specifically, spines within 5 microns (five one-millionths of a meter) acted in concert. But then, right outside that 5-micron boundary, spines were less likely than chance to join in that activity. Sur speculates that these isolated pockets of activity sharpened the response from each enclave.
“Apical” vs. “basal:” The neurons the team studied have two distinct kinds of dendrites. Apical dendrites, which are very long and protrude from the top, or “apex,” of the neuron, tend to get a wide variety of inputs from across the cortex. Basal dendrites, which are shorter and extend out from the bottom, typically get more raw visual input. While basal dendrites indeed received more visual input than apical dendrites overall, Jenks found that apical dendrites on visually responsive neurons had significantly more visually responsive spines than those on non-responsive neurons. And both types of dendrites equally obeyed the rules above about distance from the soma.
Orientation selectivity matters most: Jenks, Sur, and the team used statistical modeling to determine which of many factors (the stimulus selectivity, reliability of the response, a spine’s distance from the soma, apical versus basal, etc.) most explained how correlated a spine’s responsiveness was with that of the soma. By a wide margin, how selective a spine was to the orientation of its preferred grating was the most important single factor.
“Our results reveal that synaptic inputs to excitatory layer 2/3 neurons in mouse (visual cortex) are not randomly arranged, but organized and distributed in a manner that correlates with multiple factors including somatic responsiveness, somatic tuning, branch type, distance from the soma, local correlations, and stimulus selectivity,” the researchers wrote.
The team’s findings can help advance studies of vision in the brain in multiple ways, Jenks and Sur say. Certain genetic mutations that affect how neurons connect in circuits can affect visual cortex neurons and vision, Sur says. Documenting these rules provides researchers with a baseline to compare against when examining the effects of such mutations. Jenks adds that the findings could inform efforts to model how neurons integrate synaptic inputs in their computations.
In addition to Sur and Jenks, the paper’s other authors are Gregg Heller, Katya Tsimring, Kendyll Martin, Asrah Rizvi, and Jacque Pak Kan Ip.
The National Institutes of Health, the Simons Foundation Autism Research Initiative, and the Freedom Together Foundation provided support for the study.
MIT affiliates elected to National Academy of Sciences for 2026Six MIT faculty, along with 10 additional alumni, are recognized by their peers for their outstanding contributions to research in the natural and social sciences.The National Academy of Sciences (NAS) has elected 120 members and 25 international members for 2026, including six MIT faculty members and 10 additional alumni.
Among MIT professors, Bengt Holmström, Michale Fee, Gareth McKinley ’91, Keith Nelson, Fan Wang, and Catherine Wolfram ’96 were elected in recognition of their “distinguished and continuing achievements in original research.”
Additional alumni who were elected include Christopher J. Chang PhD ’02 (Chemistry); Cynthia J. Ebinger SM ’86, PhD ’88 (Earth, Atmospheric and Planetary Sciences); Andrew Gelman ’85, ’86 (Mathematics and Physics); Richard L. Greene ’60 (Physics); Chuan He PhD ’00 (Chemistry); Pardis C. Sabeti ’97 (Biology/Life Sciences); Robert J. Shiller SM ’68, PhD ’72 (Economics); Daniel M. Sigman PhD ’97 (EAPS); Eero Simoncelli SM ’88, PhD ’93 (Electrical Engineering and Computer Science); and Salil P. Vadhan PhD ’99 (Mathematics).
Membership in the National Academy of Sciences is one of the highest honors a scientist can receive in their career. The NAS is a private, nonprofit institution that was established under a congressional charter signed by President Abraham Lincoln in 1863. It recognizes achievement in science by election to membership, and — with the National Academy of Engineering and the National Academy of Medicine — provides science, engineering, and health policy advice to the federal government and other organizations.
Bengt Holmström is the Paul A. Samuelson Professor of Economics, emeritus. He received his doctoral degree from the Stanford Graduate School of Business in 1978 and held faculty positions at Northwestern University and Yale University before joining the MIT faculty in 1994 with a joint appointment in economics and management.
Holmström is best known for his foundational research on the theory of contracting and incentives, for which he received the 2016 Sveriges Riksbank Prize in Economic Sciences in Memory of Alfred Nobel (together with Oliver Hart of Harvard University). His extensive contributions to contract theory as applied to the theory of the firm, corporate governance, and liquidity problems in financial crises have had wide-ranging impacts, while bringing contract theory into mainstream economic thought.
In addition to the Nobel, Holmström’s research has been recognized with the Stephen A. Ross Prize in Financial Economics and the Grand Cross of the Order of the Lion of Finland. He is a member of the American Academy of Arts and Sciences, the Econometric Society, and the American Finance Association. Holmström is also an elected foreign member of the Royal Swedish Academy of Sciences and a member of the Finnish Academy of Sciences and Letters.
Michale S. Fee is the Glen V. and Phyllis F. Dorflinger Professor of Neuroscience, head of the MIT Department of Brain and Cognitive Sciences (BCS), and investigator at the McGovern Institute for Brain Research. His research explores how the brain learns and generates complex sequential behaviors. Using the zebra finch as a model system, Fee investigates the neural mechanisms underlying birdsong — a behavior that young birds learn from their fathers through trial and error, much as human infants learn to speak through babbling. His research extends far beyond birdsong — the neural circuits controlling birdsong learning are closely related to human brain circuits disrupted in Parkinson’s and Huntington’s diseases. Insights from Fee’s research could reveal new clues to the causes and potential treatments of these complex brain disorders.
After receiving his BE with honors in engineering physics at the University of Michigan in 1985, Fee studied applied physics at Stanford University, where he carried out his PhD thesis work in the laboratory of Steven Chu. In 1992, he began working as a postdoc in David Kleinfeld’s lab in the Biological Computation Research Department at Bell Laboratories. Four years later, he became a permanent member of the technical staff at Bell Labs and began working on the mechanisms of vocal sequence generation in the songbird. In 2003, he became an investigator at the McGovern Institute and a faculty member in BCS. In 2021, he was appointed BCS department head, continuing the department’s tradition of being led by scientists whose exemplary work makes MIT a world leader in brain science. Fee is a member of the American Academy of Arts and Sciences and a recipient of multiple undergraduate and graduate teaching awards at MIT.
Gareth H. McKinley ’91 is the School of Engineering Professor of Teaching Innovation in the Department of Mechanical Engineering at MIT, former associate head and interim head of the department, and co-founder of Cambridge Polymer Group. McKinley’s research interests include non-Newtonian fluid dynamics, microfluidics, extensional rheology, field-responsive materials, super-hydrophobicity, drag reduction, and the wetting of nanostructured surfaces. His work focuses on understanding the rheology of complex fluids such as surfactants, biomaterials, gels, and polymers, which are ubiquitous in foods and consumer products.
McKinley has made outstanding contributions to viscoelastic fluid mechanics, understanding flow instabilities and stretching flows. His research group has developed novel instrumentation and customized rheological analysis techniques that have driven the field of rheology for complex and soft fluids. His instrumentation and testing algorithms, along with freely-distributed code for analyzing large amplitude oscillatory shear flow, and broad-band “chirp” rheometry, are used worldwide in industry and academia .
McKinley is the author of over 390 technical publications. He has won the Publication Award of the Society of Rheology twice (2007 and 2022), as well as the 2021 Walters Award from J. Non-Newtonian Fluid Mechanics. He was awarded the Bingham Medal of The Society of Rheology in 2013, the Gold Medal from the British Society of Rheology in 2014, and the G.I. Taylor Medal from the Society for Engineering Science in 2022. In 2019, he was elected to the National Academy of Engineering and was also inducted as a fellow of the Royal Society of London. In 2023, he was awarded an honorary doctorate from the Katholieke University of Leuven, and in 2024 became a corresponding member of the Australian Academy of Sciences. In 2025, he was elected to the American Academy of Arts and Sciences and also became a foreign fellow of the Indian National Academy of Engineering.
Keith A. Nelson, the Haslam and Dewey Professor of Chemistry, earned his BS in chemistry from Stanford University. After completing his doctoral studies in physical chemistry, also at Stanford, he conducted postdoctoral research with John P. McTague at the University of California at Los Angeles. In 1982, Nelson joined the MIT Department of Chemistry as an assistant professor.
His distinguished career has been recognized with numerous honors, including the William F. Meggers Award, the Bomem-Michelson Award, and the Frank Isakson Prize for Optical Effects in Solids. Research in the Nelson Group focuses on the time-resolved optical study and control of collective transformations in condensed matter, using pulses of light in the THz, optical, and X-ray spectral ranges and laser-generated strain waves to drive the modes of motion through which these changes occur.
Fan Wang is a professor of Brain and Cognitive Sciences, investigator at the McGovern Institute, and co-director of the K. Lisa Yang and Hock E. Tan Center for Molecular Therapeutics at MIT. She investigates the neural circuits that govern the dynamic interactions between brain and body, exploring how the brain generates sensory perceptions and controls movement. Wang uses cutting-edge techniques including optogenetics, in vivo electrophysiology, and in vivo imaging to make discoveries with profound clinical implications.
By developing innovative tools to study how brain circuits work, Wang discovered distinct populations of neurons activated by anesthesia that can suppress pain without blocking sensation, and can calm anxiety by regulating automatic body functions like heart rate. She also identified the brain circuits controlling rhythmic movements essential for exploration and communication. Together, these findings reveal how emotion, physiology, movement, and consciousness are deeply interconnected.
Before coming to MIT, Wang obtained her PhD from Columbia University working with Richard Axel, and received her postdoctoral training at the University of California at San Francisco and Stanford University with Marc Tessier-Lavigne. She became a faculty member at Duke University in 2003, where she was later appointed Morris N. Broad Professor of Neurobiology. Wang became an investigator at the McGovern Institute and a faculty member in the Department of Brain and Cognitive Sciences at MIT in 2021. She is a member of the American Academy of Arts and Sciences and a recipient of multiple undergraduate teaching and graduate mentorship awards at MIT.
Catherine D. Wolfram ’96 is the William Barton Rogers Professor in Energy and professor of applied economics in the MIT Sloan School of Management. Before coming to MIT Sloan, Wolfram previously served as the Cora Jane Flood Professor of Business Administration at the Haas School of Business at the University of California at Berkeley. From March 2021 to October 2022, she served as the deputy assistant secretary for climate and energy economics at the U.S. Treasury, while on leave from UC Berkeley. Before leaving for government service, she was the program director of the National Bureau of Economic Research’s Environment and Energy Economics Program and a research affiliate at the Energy Institute at Haas. Before joining the faculty at UC Berkeley, she was an assistant professor of economics at Harvard University. She received a PhD in economics from MIT in 1996 and an BA from Harvard in 1989.
Wolfram has published extensively on the economics of energy markets. Her work has analyzed rural electrification programs in the developing world, energy efficiency programs in the United States, the effects of environmental regulation on energy markets, and the impact of privatization and market restructuring in the United States and United Kingdom. She is currently working on projects at the intersection of climate, energy, and trade, including work on carbon border adjustment mechanisms and oil market sanctions. Since March 2025, Wolfram has served on the COP30 President’s Council on Economics, Finance, and Climate, and has chaired a working group on climate coalitions.
Four from MIT named 2026 Searle ScholarsComputational neuroscientist Sven Dorkenwald and cell biologist Whitney Henry, along with two MIT alumni, are recognized for their exceptional early-career research contributions.MIT scientists Sven Dorkenwald and Whitney Henry have been named 2026 Searle Scholars, an award given annually to 15 exceptional early-career researchers in the fields of biomedical sciences and chemistry. Dorkenwald is an assistant professor of brain and cognitive sciences and an investigator at the McGovern Institute for Brain Research. Henry is the Robert A. Swanson (1969) Career Development Professor of Life Sciences and an intramural faculty member at the Koch Institute for Integrative Cancer Research.
In addition, MIT alumni Irene Kaplow ’10 and Jared Mayers PhD ’15 were also honored.
Chosen by a scientific advisory board, Searle Scholars are considered among the most creative young researchers pursuing high-risk/high-reward research. The Searle Scholars Program is funded through the Searle Funds at The Chicago Community Trust and administered by Kinship Foundation. Each scholar will each receive $450,000 in flexible funding to support their work over the next three years.
Sven Dorkenwald
Sven Dorkenwald is a computational neuroscientist investigating the organizational principles of neuronal circuits. The synaptic connectivity of neurons, their connectome, is fundamental to how networks of neurons function. Dorkenwald develops computational and collaborative tools to map, analyze, and interpret synapse-resolution connectomes. His work has led to large connectomic reconstructions of the fruit fly brain and parts of mammalian brains. He uses these connectomes to investigate the architecture of neuronal circuits and how their structure supports complex computations.
“As I establish my new lab, the Searle Scholars Award will help us launch ambitious projects and set our long-term scientific direction,” says Dorkenwald. “I am deeply grateful for the support from the Kinship Foundation and look forward to interacting with this amazing cohort of Searle Scholars.”
Dorkenwald joined the faculty of MIT in 2026 as an assistant professor in the Department of Brain and Cognitive Sciences and an investigator at the McGovern Institute. He earned a BS in physics and an MS in computer engineering from the University of Heidelberg, followed by a PhD in computer science and neuroscience at Princeton University in 2023 under the mentorship of Sebastian Seung and Mala Murthy. Dorkenwald completed his postdoctoral training as a Shanahan Research Fellow at the Allen Institute and the University of Washington, while serving as a visiting faculty researcher at Google Research.
Whitney Henry
Whitney Henry investigates the potential of ferroptosis, an iron-dependent form of cell death, for developing novel therapies that target subpopulations of cancer cells that are highly metastatic, therapy-resistant, and therefore critical instigators of tumor relapse. Her research is focused on uncovering the molecular factors influencing ferroptosis susceptibility, investigating its effects on the tumor microenvironment, and developing innovative methods to manipulate ferroptosis resistance in living organisms, drawing from functional genomics, metabolomics, bioengineering, and a range of in vitro and in vivo models.
“I am incredibly grateful to the Kinship Foundation for supporting our research and giving us the freedom to ask bold, curiosity-driven scientific questions,” says Henry. “This support allows us to pursue ambitious ideas, take creative risks, and embark on new research directions.”
Henry joined the MIT faculty in 2024 as an assistant professor in the Department of Biology and a member of the Koch Institute, and is currently an HHMI Freeman Hrabowski Scholar. She received her bachelor's degree in biology with a minor in chemistry from Grambling State University and her PhD from Harvard University. Following her doctoral studies, she worked in the lab of Robert Weinberg at the Whitehead Institute for Biomedical Research and was supported by fellowships from the Jane Coffin Childs Memorial Fund for Medical Research and the Ludwig Center at MIT.
Alumni also honored
Irene Kaplow ’10, a graduate of the MIT Department of Mathematics, is an assistant professor in the Department of Biology and the Ray and Stephanie Lane Computational Biology Department at Carnegie Mellon University. Her selection as a Searle Scholar is for “deciphering transcriptional regulatory mechanisms underlying mammalian dietary phenotype evolution and their relationships to transcriptional regulatory responses to changes in diet.”
Jared Mayers PhD ’15, who earned his doctorate from the MIT Department of Biology, is an assistant professor at the Fred Hutchinson Cancer Center at the University of Washington. His selection as a Searle Scholar is for “a reverse-translational framework to decipher metabolic vulnerabilities of bacterial pathogens.”
The Haystack 37m Telescope: A new era of astrophysical researchThe legendary radio astronomy telescope returns to its science and educational mission at MIT Haystack Observatory.The Haystack 37m Telescope has been a landmark in radio astronomy and radar studies of the solar system since its first light in 1964. Over the following four decades, it supported NASA's Apollo landings on the moon, made planetary radar maps of the surface of Venus, contributed to experimental tests of Einstein's general relativity, supported the development of VLBI, and conducted foundational studies of quasars and star-forming regions.
Recently, the Haystack 37m Telescope — a 37-meter radio and millimeter-wavelength antenna at MIT Haystack Observatory in Westford, Massachusetts — made its return to front-line astronomical research following an extended period of system upgrades. These observations reconnect this instrument with its long tradition of scientific discovery and open a new chapter.
On Dec. 8, 2025, Haystack scientists observed the supermassive black hole system at the center of the galaxy Messier 87 (M87) using a technique called very long baseline interferometry (VLBI) that links telescopes across continents to achieve extraordinary resolution. These observations mark the return of one of America's most storied radio telescopes to its historical scientific and educational mission.
The observations targeted the powerful jet of energy and matter launched from M87’s central black hole, M87*. This jet, driven by a black hole six-and-a-half billion times the mass of our sun, extends thousands of light years into intergalactic space and is one of the most energetic phenomena in the known universe.
Previous international campaigns, namely those led by the Event Horizon Telescope, have imaged the black hole's immediate “shadow.” The Haystack 37m Telescope observations, performed in collaboration with the telescopes of the Very Long Baseline Array (VLBA) and the Greenland Telescope (GLT), help to probe the larger-scale structure of the jet, investigating how energy is transported far beyond the black hole's vicinity. Understanding this process is central to explaining how supermassive black holes shape the galaxies that surround them.
“The Haystack 37m Telescope’s exceptional sensitivity enables the intercontinental telescope array to detect faint emission from around the distant M87* black hole,” says Paul Tiede, principal investigator of the M87 study. “In tandem with the GLT and the VLBA, Haystack is helping create the first multifrequency movies of M87*’s faint jet, greatly improving our understanding of black hole physics.”
The upgraded Haystack 37m Telescope opens multiple new lines of research. At MIT, Saverio Cambioni and Richard Teague of the Department of Earth, Atmospheric and Planetary Sciences (EAPS) plan to use the instrument within MIT’s Planetary Defense Project to measure asteroid sizes and shapes, characterizing objects that could pose a hazard to Earth and deepening our understanding of the solar system’s formation. Associate Professor Brett McGuire of the Department of Chemistry plans to search for complex organic molecules in space, work that speaks to the question of how the chemical precursors to life arise.
“We are thrilled to provide the research community with a powerful telescope at a time where few such instruments are available,” says Jens Kauffmann, principal investigator of the Haystack 37m Telescope Astronomy Program, who uses the telescope to study the formation of stars and their planets. “Even more exciting are the prospects this generates for the next generation of astronomers. Hands-on training opportunities on world-class research telescopes have become exceptionally rare worldwide, and now we can offer this singular advanced workforce development program right here in Massachusetts.”
Student involvement with the Haystack 37m Telescope has already resumed: Undergraduate interns at Haystack Observatory played an active role in developing the telescope’s control systems and data analysis algorithms. This work exemplifies Haystack’s role as a hands-on research and training environment where students contribute directly and gain practical experience with a frontline research instrument.
The return to research-focused observations is the result of more than 10 years of careful, sustained work. From 2010 to 2014, the Haystack 37m Telescope underwent a major upgrade and refurbishment that enhanced its ability to observe at millimeter wavelengths. This work was primarily done to improve the antenna’s capability as a space radar. The dish now primarily serves U.S. government agencies in that capability, and astronomy was temporarily a secondary activity.
But work to restore the telescope's science capability never stopped. Initial support from the National Science Foundation (NSF) in 2015 modernized systems for data analysis and radio signal processing. The first successful engineering-oriented VLBI experiments with the new dish were conducted at the same time. Additional NSF funding in 2019, provided in the context of the Next Generation Event Horizon Telescope (ngEHT) program, enabled a more general and sustained effort to upgrade receiver equipment and computing systems. Support from private donors to Haystack also aided in this longer-term effort.
Several recent developments, particularly in 2025, proved significant. With support from MIT's Jarve Seed Fund for Science Innovation, scientists and engineers removed lingering technical limitations with astronomy systems and expanded the telescope's scientific reach. Other funding for projects led by the Smithsonian Astrophysical Observatory enabled the M87 campaign and commissioning of the next-generation digital back end, a highly advanced signal-processing system developed for the ngEHT. Together, these advances made the December 2025 observations possible. MIT Haystack Observatory is now pursuing support from both private and federal sources for further improvements under the Haystack 37m Telescope Astronomy Program.
“The upgraded Haystack 37m Telescope empowers MIT students and researchers to pursue fundamental questions relating to our origins and our solar system,” says Richard Teague, professor at MIT EAPS. “With privileged access to such a powerful facility, we can undertake ambitious observational programs previously impossible to schedule. This is the beginning of what we expect will be an exciting era of new discoveries with the Haystack 37m Telescope.”
Single-molecule tracker illuminates workings of cancer-related proteinsResearchers can now use custom-built microscopy and nanotechnology to tag and follow the activity of individual proteins in real-time.Using a powerful single-molecule imaging method they developed, a research team from the Broad Institute of MIT and Harvard has unveiled a dynamic view of how some cancer-related proteins interact in living cells.
The technique relies on highly stable nanoparticle probes that brightly illuminate individual molecules for long periods of time. The researchers used their method to observe, for the first time, individual receptors as they move around the cell membrane, attaching to and then letting go of other receptors to alter signaling within the cell.
Described in the journal Cell, the work demonstrates the method’s potential for investigating other receptors and molecules, and for improved drug screening to better understand the effects of therapeutics on living cells.
“With our photostable probes, we can map out the entire lifespan of these molecules in their native environment and see things that have never been observable before,” says study leader Sam Peng, a Broad Institute core institute member and assistant professor of chemistry at MIT.
Molecular movies
Peng’s method solves a problem with existing contrast agents used in single-molecule tracking, such as dyes. Under the laser light that’s used to excite these dyes, they burn out after a few seconds in a phenomenon known as photobleaching, which means that scientists could only use them to take a few snapshots of cell receptors, and not follow them over the entirety of the signaling process.
For a longer and richer view, Peng’s lab developed long-lasting probes, known as upconverting nanoparticles, which emit signals that remain stable under laser excitation. The nanoparticles contain rare-earth ions that continue to luminescence for minutes, hours, and potentially years. In addition, by altering the type and doses of the ions, scientists can engineer probes emitting in many different colors, enabling tracking of many targets in a single experiment.
In the current study, the researchers aimed to uncover new biology by focusing on the EGFR family of cell receptors, which have been linked to several kinds of cancer. They collaborated with EGFR experts Matthew Meyerson and Heidi Greulich of the Broad’s Cancer Program. They knew that EGFR receptors need to pair up, or “dimerize,” in order to initiate signaling within the cell, but they wanted to learn more about the dynamics of these pairings — what the receptors partner with, how long they stay together, and how they find new partners.
For a better and more sustained look at the receptors, the research team customized their upconverting nanoparticles to tag EGFR and related receptors HER2 and HER3, which are linked to cancer, and used them to track the molecules in living human cells.
A new view of protein pairings
In this study, Peng and his team observed that, when activated with a stimulating molecule, EGFR receptors can pair up and stay dimerized for several minutes, something not observable using traditional dyes. Excessive and prolonged dimerization can lead to too much cell growth and cancer.
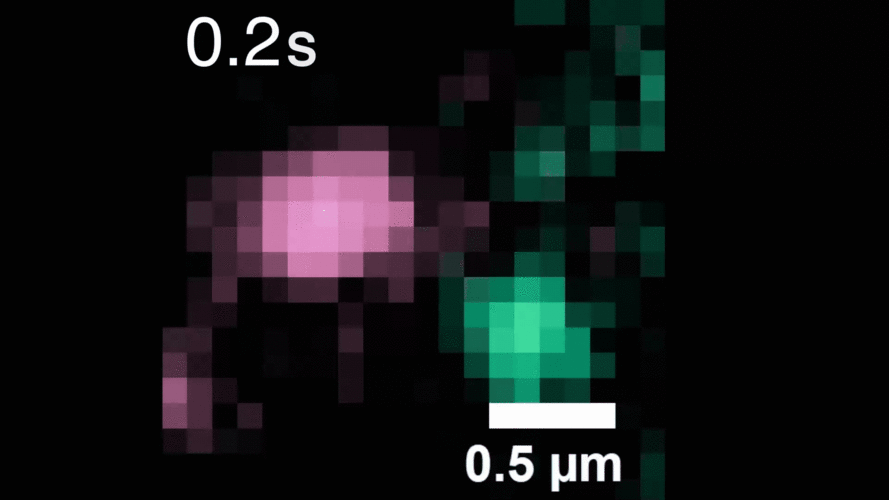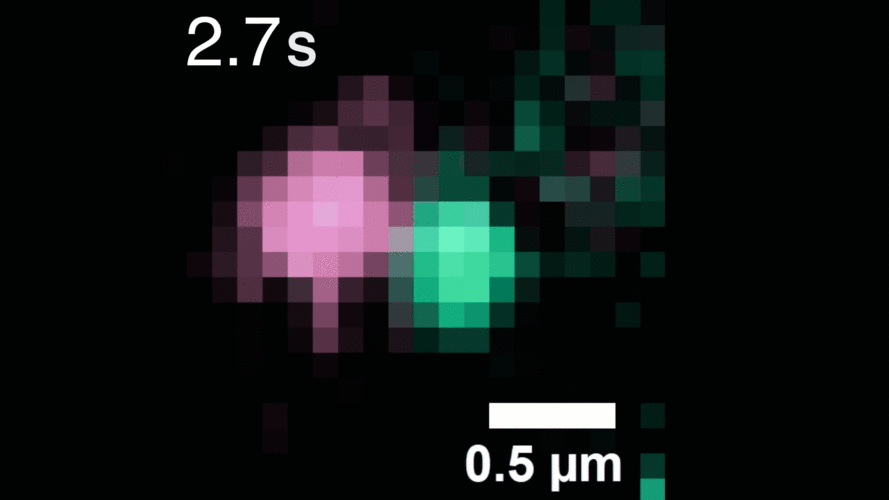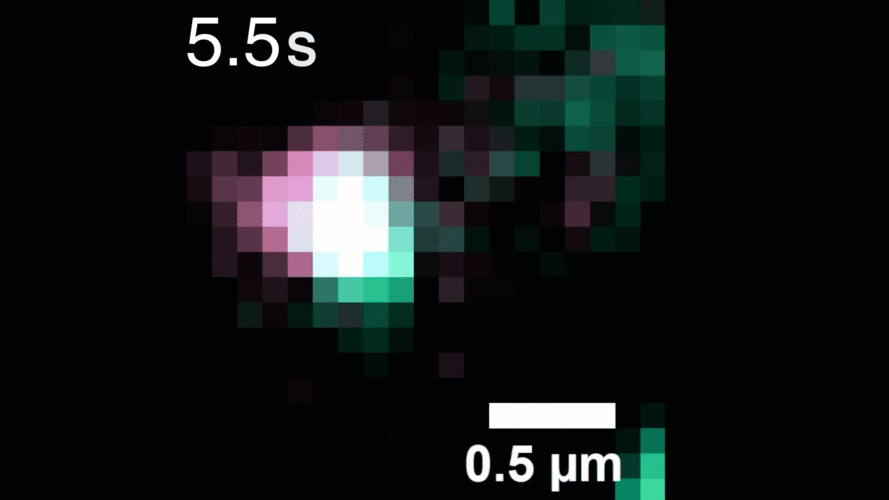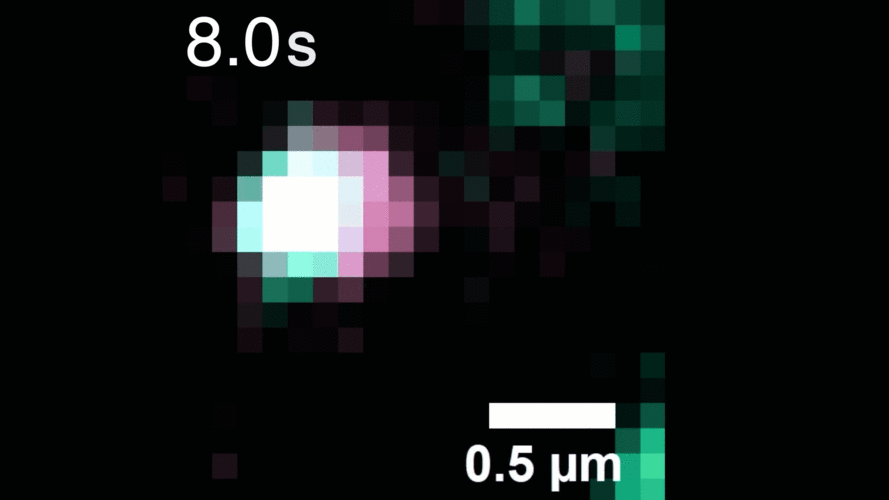 A microscopy video shows upconverting nanoparticles tagged to EGFR receptors (labeled pink and green), which track individual receptors as they dimerize. Image courtesy of the researchers.
A microscopy video shows upconverting nanoparticles tagged to EGFR receptors (labeled pink and green), which track individual receptors as they dimerize. Image courtesy of the researchers.
When the EGFR molecules carried cancer-related mutations, the dimers became more stable, with the more stabilizing mutations linked to more potent cancers in people. In addition, the mutated receptors could form stable dimers even without an external stimulus prompting them to dimerize. The finding helps explain how EGFR mutations can lead to uncontrolled cell growth and cancer, and could inform efforts to target this process therapeutically.
The team discovered several other new and surprising details about how HER2 and HER3 form stable pairings with themselves, which helps illuminate the role of these molecules in related cancers.
When the research team tagged all three receptor types in one experiment, they observed a vibrant scene with receptors navigating the cell surface, finding partners, unpairing, and then finding new partners, over and over again.
Beyond shedding light on EGFR biology, the scientists hope that collaborators in other fields will apply their method to ask new scientific questions about other proteins of interest. “We think this technique could be transformative for studying molecular biology, because it enables dynamic biological processes to be observed with high spatiotemporal resolution over unprecedented timescales,” says Peng.
They are also planning to explore the method’s use in studying the mechanism of drug action, to reveal how potential therapeutics alter individual molecules over time. In addition, they will continue to improve their methods, such as making the probes smaller, brighter, and able to emit more colors.
Language development in the brainThe brain’s language network is still evolving in adolescence. But by age 4, language processing is already handled by the left side of the brain, new research finds.The brain’s capacity to use and understand language expands rapidly in the first years of life, as babies start to make sense of the words they hear and eventually begin to piece together sentences of their own. The language-processing parts of the brain that make this possible continue to evolve in older children, as they expand their vocabularies and learn to use language more flexibly.
MIT brain researchers have captured snapshots of the developing language-processing network in brain scans of hundreds of children and adolescents. Their data, reported May 16 in the journal Nature Communications, show that the network continues to mature, becoming better integrated and increasingly responsive until around age 16. But they also found that a key feature of the adult language network is established early on: its localization in the left side of the brain.
Language lateralization
It is well known that using language is mostly the job of the left hemisphere. As adults, we call on the language-processing regions there when we read, write, speak, or listen to others talk. But there was some question as to whether this left lateralization is established early in life, or might instead emerge as the language network matures, with both sides of the brain contributing to language in childhood.
To find out, researchers needed to see young brains in action — and several MIT labs had collected exactly the right kind of data. Groups led by Evelina Fedorenko, an associate professor of brain and cognitive sciences; John Gabrieli, the Grover Hermann Professor of Health Sciences and Technology; and Rebecca Saxe, the John W. Jarve (1978) Professor of Brain and Cognitive Sciences, teamed up to share brain scans from children, adolescents, and adults and compare how their brains responded to language. Fedorenko, Gabrieli, and Saxe are also investigators at the McGovern Institute for Brain Research.
In studies aimed at better understanding a variety of cognitive functions and developmental disorders, the three teams had all collected functional MRI data while subjects participated in “language localizer” tasks — an approach the Fedorenko lab developed to map the language-processing network in a person’s brain. By monitoring brain activity with functional MRI as people engage in both language tasks and non-linguistic tasks, researchers can identify parts of the brain that are exclusively dedicated to language processing, whose precise anatomic location varies across individuals.
To activate the language network, the researchers had children listen to stories inside the MRI scanner. Depending on their age, some heard excerpts of “Alice in Wonderland,” some listened to podcasts and TED talks, and others heard shorter, simpler stories. To watch their brains during a non-linguistic task, the researchers had the children listen to nonsense words.
Across the data from the three labs, which included children between the ages of 4 and 16, as well as adults for comparison, the team saw clear developmental changes in the brain’s response to language. “The integration of the system — how well different subregions of the system correlated with each other and worked together during language processing — was stronger in older children as compared to younger children,” says Ola Ozernov-Palchik, a research scientist in Gabrieli’s lab and a research assistant professor at Boston University. The system was also more strongly activated by language in older children, which may reflect their growing comprehension of what they hear.
But strikingly, almost all language processing happened on the left side of the brain, even in the youngest subjects. “From age 4 on, it seems just as lateralized as in an adult,” Gabrieli says.
Language and developmental disorders
The researchers say this finding has implications for understanding developmental conditions that impact language, including autism and dyslexia. The right side of the brain frequently gets more involved in language processing in people with these conditions than it does in typically developing children. “Almost every single developmental disorder that’s associated with language has a theory that’s related to language lateralization,” Ozernov-Palchik says.
The reason for more bilateral language processing in some disorders is debated. One idea has been that some people might use both sides of their brain for language processing because their brains are less mature. If the right side of the brain processes language early in life, scientists had reasoned, it might simply continue to do so for longer in people with autism or dyslexia than it does in neurotypical individuals. But if most people use the left side of their brains for language even when they are young, the difference can’t be attributed to a developmental delay. Other developmental differences might cause bilateral language processing instead.
The researchers don’t have the full picture yet; they still need to know what parts of the brain process language in children younger than 4. Likewise, they would like to know what the brain areas that become the language network are doing in the first months of life, when infants aren’t using language yet. They are eager to find out, both to understand fundamentals of brain development and to shed light on developmental disorders. “I think understanding that normal trajectory is really critical for interpreting what a deviation from that trajectory is,” says Amanda O’Brien, a former graduate student in Gabrieli’s lab who is now a postdoc at Harvard University.
One reason people thought lateralization might develop gradually is because damage to the left hemisphere of the brain impacts language abilities differently, depending on when it occurs. “If you have damage to the left hemisphere as an adult, you’re very likely to end up with some form of aphasia, at least temporarily,” Fedorenko explains. “But a lot of the time, with early damage to the left hemisphere, you grow up and you’re totally fine. The language can just develop in the right hemisphere.”
Some scientists suspected that the right side of the brain was able to take over language processing in children who suffered early-life brain damage because it was already participating in this function at the time. But the team’s findings suggest the developing brain may be nimbler than that. “Our data tell you that this early plasticity apparently happens in spite of the fact that by age 4, we see these very strongly lateralized responses already,” Fedorenko says.
Two from MIT named 2026 Knight-Hennessy ScholarsThe prestigious fellowship funds graduate studies at Stanford University.MIT master’s student Sunshine Jiang ’25 and Rupert Li ’24 are recipients of this year’s Knight-Hennessy Scholarship. Now in its ninth year, the highly competitive scholarship provides up to three years of financial support for graduate studies at Stanford University.
Sunshine Jiang ’25
Sunshine Jiang, from Hangzhou, China, graduated from MIT in 2025 with a bachelor’s degree as a double major in physics and electrical engineering and computer science, along with minors in mathematics and economics. She will receive her master of engineering degree this month and will start her PhD in computer science at Stanford School of Engineering this fall.
Jiang researches embodied artificial intelligence and robotics, developing data-efficient, adaptive systems for general-purpose robots that broaden accessibility. She has presented her research at major conferences, including the Conference on Robot Learning, the International Conference on Robotics and Automation, and the International Conference on Learning Representations.
Jiang led the development of AI-powered systems that provide access to traditional Chinese art in rural classrooms, founded cross-country programs that expand girls’ access to STEM education, and created a Covid-19 documentary amplifying community voices, which was featured on China Daily.
Rupert Li ’24
Rupert Li, from Portland, Oregon, is currently pursuing a PhD in mathematics at Stanford School of Humanities and Sciences. He graduated from MIT in 2024 with a bachelor’s degree, double majoring in mathematics and computer science, economics, and data science. Along with his bachelor’s degree, he also received a master’s degree in data science. Li then traveled to the United Kingdom as a Marshall Scholar, where he earned a master’s degree in mathematics from the University of Cambridge.
Li’s research interests lie in probability, discrete geometry, and combinatorics. He enjoys serving as a mentor for MIT PRIMES-USA, a high school math research program, and previously served as an advisor for the Duluth REU, an undergraduate math research program. In addition to the Knight-Hennessy Scholarship and the Marshall Scholarship, he has been awarded the Hertz Fellowship, P.D. Soros Fellowship, and the Goldwater Scholarship, and he received honorable mention for the Frank and Brennie Morgan Prize.
MIT chemists discover and isolate a new boron-oxygen moleculeThe discovery of dioxaborirane could expand the chemistry of boron-based reagents, providing new tools for oxidation reactions in synthesis and materials science.Oxygen is a cornerstone of chemistry, largely because it is so good at building the organic molecules that make up our world. Some oxygen-based compounds, called peroxides, are famous for being highly reactive — they act like oxygen delivery trucks, transferring atoms to other molecules. This process is essential for everything from creating new medicines to industrial manufacturing.
In an open-access study published April 24 in Nature Chemistry, researchers from the labs of MIT professors Christopher C. Cummins and Robert J. Gilliard, Jr. have revealed a brand-new type of peroxide containing boron. This molecule, called a dioxaborirane, represents a major advance in a field where such structures were long-proposed, but considered too unstable to actually isolate.
Room-temperature breakthrough
Dioxaborirane forms when a specially engineered boron molecule reacts with oxygen gas. What makes this discovery remarkable is that the reaction happens almost instantly at room temperature. Usually, creating strained oxygen-containing rings like this requires extreme, “punishing” conditions — like freezing temperatures or high pressure — to keep the molecule from falling apart.
Using advanced tools such as crystallography and computational modeling, the team proved the existence of a highly strained, three-member ring made of one boron and two oxygen atoms.
A molecule with two personalities
The most exciting part of the discovery is how the molecule behaves. Depending on its electrical charge, it acts in two very different ways:
“By showing that these compounds can be generated under mild conditions, our work opens the door to entirely new types of chemistry,” says Chonghe Zhang, the first author of the paper and an MIT chemistry graduate student co-advised by Cummins and Gilliard. “In the long term, these findings could provide us with powerful new tools for oxidation reactions in synthesis and materials science.”
Additional co-authors on the paper are Noah D. McMillion and Chun-Lin Deng of MIT and Junyi Wang of Baylor University. The work was funded, in part, by the U.S. National Science Foundation.
A new way to spot signs of dark matterGravitational waves emitted by colliding black holes may bear imprints of dark matter, which physicists could detect with a new model.Dark matter is thought to make up most of the matter in the universe, but the only way it interacts with its surroundings is through gravity. If two colliding black holes spiral through a dense region of dark matter and merge, gravitational waves rippling across space and time could carry an imprint of that dark matter.
Now, physicists may be able to spot such imprints of dark matter in gravitational waves that are detected on Earth.
Researchers at MIT and in Europe have developed a method that makes predictions for what a gravitational wave should look like if it were produced by black holes that moved through dark matter, rather than empty space. They applied the technique to publicly available gravitational-wave data previously recorded by LIGO-Virgo-KAGRA (LVK), the global network of observatories that detect gravitational waves from black hole mergers and other far-off astrophysical sources.
The researchers looked through the gravitational-wave signals recorded over the LVK’s first three observing runs. From 28 of the clearest signals, the team found that 27 originated from black holes that merged in a vacuum, as physicists expected. But the pattern of one signal, GW190728, showed possible signs of a dark matter imprint.
The scientists emphasize that they have not detected dark matter. Rather, the new method offers a new way to screen gravitational-wave data for hints of dark matter, which physicists can then follow up and confirm with other techniques.
“We know that dark matter is around us. It just has to be dense enough for us to see its effects,” says Josu Aurrekoetxea, a postdoc in the MIT Department of Physics. “Black holes provide a mechanism to enhance this density, which we can now search for by analyzing the gravitational waves emitted when they merge.”
Aurrekoetxea and his colleagues report their results in a study appearing today in Physical Review Letters. The study’s co-authors are LVK member Soumen Roy of Université Catholique de Louvain (UCLouvain) in Belgium, Rodrigo Vicente of the University of Amsterdam, Katy Clough of Queen Mary University of London, and Pedro Ferreira of Oxford University.
A dark pull
Dark matter is an invisible, hypothetical form of matter that, unlike normal everyday matter, has no interactions with the electromagnetic force. Dark matter can pass through light, magnetic fields, and any other form of energy along the electromagnetic spectrum without leaving a trace. The only evidence that dark matter exists is through its apparent interaction with one other force: gravity.
By observing how gravity bends around distant galaxies, astronomers have surmised that there must be an extra force, outside of the galaxies’ own gravitational pull, to explain the bending fields, or “lensing.” This extra force, physicists suspect, is dark matter, which could account for over 85 percent of the matter in the universe. But exactly what dark matter is is a matter of huge debate, with theories for dark matter particles that range widely in particle size and properties.
One class of proposed dark matter consists of “light scalar” particles, whose masses are many orders of magnitude lighter than an electron. Theorists predict that such dark matter should behave not just as particles, but also as coordinated waves when moving near black holes.
When waves of dark matter come in contact with a rapidly spinning black hole, physicists predict that the black hole's rotational energy can be transferred to the dark matter, amplifying it. This phenomenon, known as superradiance, would whip up the waves to extremely high densities of dark matter, akin to churning cream into butter.
At high enough densities, light scalar dark matter, which is invisible by all other accounts, should leave an imprint on the gravitational waves that reverberate from the colliding black holes.
But exactly what would that imprint look like? And could such an imprint be detectable in gravitational waves that arrive on Earth, from black holes that merged many millions of light years away?
For answers to those questions, Aurrekoetxea and his colleagues developed a model to predict the gravitational waveform, or the pattern of gravitational waves that two black holes would produce, if they collided in an environment of dark matter, versus in a vacuum (empty space, with no dark matter).
An imprint’s prediction
For their new study, the team performed detailed numerical simulations to predict the gravitational wave that would be produced given various properties of two colliding black holes — a system known as a “black hole binary.” They considered black hole binaries across a range of scenarios and properties, for example, varying the size and mass of each black hole, the environment of dark matter that the black holes might pass through, and the density of the dark matter that the black holes would spin up.
They designed the model to predict what a gravitational wave from a black hole binary would look like if it carried an imprint of dark matter, and furthermore, what that wave would look like if it traveled a given distance across space and time, to eventually arrive at a detector on Earth.
With their model, they looked to see whether any gravitational-wave signals that have been detected on Earth match their predicted patterns of dark matter imprints. To do so, they applied the model to publicly-available data recorded by LVK over the observatories’ first three observing runs. The observatories have picked up hundreds of gravitational-wave signals during this period. For their purposes, the researchers focused on the clearest signals, comprising gravitational waves from 28 separate events.
For each event, the team compared the pattern of the actual gravitational wave against their model of what the signal would look like if it were generated by the same event in an environment of dark matter. They also compared the gravitational wave to the more expected scenario in which the signal was produced in a vacuum.
Of the 28 clearest signals that they analyzed, 27 were solidly within the predictions for having been produced in a vacuum. However, the pattern of one event, GW190728, showed a “preference,” or an agreement with the team’s dark matter model. In other words, the signal may carry an imprint of dark matter.
GW190728 is a gravitational wave that is named after the date that it was detected — on July 28, 2019. Scientists previously determined that the gravitational wave originated from a black hole binary with a total mass of about 20 times the mass of the sun. With their model, the team showed that such a system could have merged through a dense cloud of dark matter and produced a similar gravitational wave to GW190728.
“The statistical significance of this is not high enough to claim a detection of dark matter, and further checks should be performed by independent groups,” Aurrekoetxea says. “What we think is important to highlight is that without waveform models like ours, we could be detecting black hole mergers in dark matter environments, but systematically classifying them as having occurred in vacuum.”
“We now have the potential to discover dark matter around black holes as the LVK detectors keep collecting data in the coming years,” says co-author Soumen Roy, who led the data analysis part of the work. “It is an exciting time to search for new physics using gravitational waves.”
“Using black holes to look for dark matter would be fantastic,” adds co-author Rodrigo Vicente, who developed the analytical model of the signal. “We would be able to probe dark matter at scales much smaller than ever before.”
This work was supported, in part, by the U.S. National Science Foundation and MIT’s Center for Theoretical Physics — a Leinweber Institute.
Powerful shrinking technique could enable devices that compute with lightMIT researchers created tiny 3D photonic devices with features small enough to channel visible light.Using a new technique that can create vacancies at any site across a material and then shrink it to about 1/2,000 of its original volume, MIT researchers have designed nanotechnology devices that could be used for optical computing and other applications involving the manipulation of visible light.
The new fabrication technique, known as “implosion carving,” allows researchers to imprint features throughout a hydrogel using photopatterning. If patterned with a resolution of about 800 nanometers, these features can then be shrunk to less than 100 nanometers.
Because that resolution is smaller than the wavelength of light, the devices can bend light in specific ways that allow them to perform optical computations.

“In order to enable nanophotonic applications in visible light, we need to make nanostructures with feature sizes with a resolution less than 100 nanometers. Only in that way can we precisely create the structure that can manipulate visible light,” says Quansan Yang, a former MIT postdoc, now an assistant professor at the University of Washington, and one of the lead authors of the new study.
In their paper, the researchers demonstrated a photonic device that can perform a simple digit-classification task, but future versions could be used for high-speed imaging and information processing, they say.
Gaojie Yang, a former MIT postdoc, is the co-lead author of the paper, which appears today in Nature Photonics. The paper’s senior authors are Peter So, director of the MIT Laser Biomedical Research Center (LBCR) and an MIT professor of biological engineering and mechanical engineering, and Edward Boyden, the Y. Eva Tan Professor in Neurotechnology at MIT and a professor of biological engineering, media arts and sciences, and brain and cognitive sciences. Boyden is also a Howard Hughes Medical Institute investigator and a member of MIT’s McGovern Institute for Brain Research, the Yang Tan Collective, and Koch Institute for Integrative Cancer Research.
Nanoscale feature sizes
Photonic devices, which transmit and manipulate light, hold potential for use as optical computer chips that could offer an energy-efficient alternative to semiconductor chips. However, existing techniques for creating 3D photonic devices haven’t yet achieved the 100-nanometer resolution that is needed to channel visible light, which has wavelengths between 380 and 750 nanometers.
Using an additive manufacturing technique called two-photon lithography, researchers can use light to create 3D nanoscale features, but with a resolution larger than 100 nanometers. Another technique, known as electron-beam lithography, can be used to etch smaller-resolution features onto a silicon chip, but it doesn’t generate 3D structures.
To make 3D devices with the necessary feature size, the researchers extended the concept of “implosion fabrication,” which Boyden’s lab developed in 2018, to create a new variant called “implosion carving.” In implosion carving, a laser creates vacancies — tiny voids where the hydrogel material has been removed — at precisely targeted locations. These vacancies exhibit different optical properties than the surrounding hydrogel. The hydrogel is then shrunk to bring the patterned features down to the nanoscale.
The carving process begins with immersing the hydrogel in a photosensitizing dye. Then, the researchers use a laser to excite the photosensitizer at specific places in the gel, which in turn generates reactive oxygen species that cut the bonds holding the hydrogel together. This creates a vacancy in that spot.
Once the desired vacancy pattern has been carved into the hydrogel, the researchers shrink it using a two-step process. First, they soak it in a solution containing ions, which causes it to shrink about tenfold in each dimension. To shrink it a little more, and to remove the watery solution, the hydrogel then undergoes a process called supercritical drying, which can remove liquid from a gel without damaging it.
At the end of the process, the hydrogel has been shrunk more than tenfold in each dimension, leading to a 2,000-fold reduction in volume.
Computing with light
To demonstrate the versatility of this technique, the researchers used it to create several 3D shapes, including a helix and a structure inspired by a butterfly wing. Some of these structures are too thin, and have too high an aspect ratio, to be stably created using conventional two-photon lithography.
The researchers also created a device that could perform a simple calculation known as digit classification, a task that is traditionally used to test the performance of neural networks. During this task, the device was presented with a digit, such as 1 or 5, and had to light up a specific location to indicate which number was detected.
To achieve this, the researchers patterned vacancies throughout the device so that it would act like a neural network. The pattern of vacancies would diffract input light as it passed through many layers of patterned hydrogel, so that the output light was determined by the shape of the digit that was entered into the system.
“This is a purely optical system that effectively performs optical computing,” So says.
“One of the very attractive features of this technology is that you can manipulate the property of the material at every tiny location,” says Dushan Wadduwage, an assistant professor at Old Dominion University and former MIT postdoc, who is also an author of the paper. “You have millions of different locations that you need to decide the property of, and that turns into a really interesting design problem where we can use deep-learning algorithms to find designs over these millions of parameters and come up with parts that go into optical systems in new ways.”
The researchers now plan to use the same principles to build optical devices that could classify cells based on their state as they flow through a microfluidic device. This could help identify rare cells such as circulating tumor cells in a blood sample, they say.
This approach could also enable the creation of high-throughput imaging techniques for applications such as analyzing tissue samples from biopsies or surgical specimens. And, if adapted to work with other materials such as hydrophobic polymers, it could also be used to create channels within 3D nanofluidic devices.
Other authors of the paper include Gaojie Yang, Takahiro Nambara, Hiroyuki Kusaka, Yuichiro Kunai, Alex Matlock, Corban Swain, Brett Pryor, Yannick Salamin, Daniel Oran, Hasindu Kariyawasam, Ramith Hettiarachchi, and Marin Soljacic.
The research was funded, in part, by the MIT-Fujikura Partnership Fund, the U.S. Army Research Office through the Institute for Soldier Nanotechnologies at MIT, Lisa Yang and Y. Eva Tan, John Doerr, the Open Philanthropy Project, the Howard Hughes Medical Institute, and the U.S. National Institutes of Health.
Improving the reliability of circuits for quantum computersA new technique helps scientists measure a phenomenon that can cause quantum circuits to perform differently than expected, increasing the error in computations.Quantum computers could someday solve pressing problems that are too convoluted for classical computers, such as modeling complex molecular interactions to streamline drug discovery and materials development.
But to build a superconducting quantum computer that is large and resilient enough for real-world applications, scientists must precisely engineer thousands of quantum circuits so they perform operations with the lowest possible error rate.
To help scientists design more predictable circuits, researchers from MIT and Lincoln Laboratory developed a technique to measure a property that can unexpectedly cause a superconducting quantum circuit to deviate from its expected behavior. Their analysis revealed the source of these distortions, known as second-order harmonic corrections, leading to underperforming circuit architectures.
The MIT researchers fabricated a device to detect second-order harmonic corrections, identify their origin, and precisely measure their strength. This technique could help scientists deliberately design quantum circuits that can counteract the effects of these deviations.
This is especially important in larger and more complicated quantum circuits, where the negative impact of second-order harmonic corrections can be amplified.
“As we make our quantum computers bigger and we want to have more precise control over the parameters of these devices, identifying and measuring these effects is going to be important for us to have a precise understanding of how these systems are constructed. It is always important to keep diving down into the circuit to see if there is an effect you didn’t expect, which impacts how your device is performing,” says Max Hays, a research scientist in the Engineering Quantum Systems (EQuS) group of the Research Laboratory of Electronics (RLE) and co-lead author of a paper on this research.
Hays is joined on the paper by co-lead author Junghyun Kim, an electrical engineering and computer science (EECS) graduate student in the EQuS group; senior author William D. Oliver, the Henry Ellis Warren (1894) Professor of EECS and professor of physics, leader of the EQuS group, director of the Center for Quantum Engineering, and associate director of RLE; as well as others at MIT and Lincoln Laboratory. The research appears today in Nature Physics.
A pair-wise problem
In a quantum computer that utilizes superconducting circuits, which is one of many potential computing platforms, Josephson junctions are critical elements that enable the transfer and manipulation of information. These devices utilize two superconducting wires that are brought very close together, with a nanometer-scale barrier between them. Like a traditional circuit, the electric charge in Josephson junctions is carried by electrons.
But in a superconducting circuit, charge-carrying electrons pair up, forming what are called Cooper pairs. These Cooper pairs can “quantum tunnel” through the barrier between the two wires, transporting current from one wire to the other.
Cooper pairs can usually only tunnel one pair at a time, which is a key property that makes quantum computation possible.
“If you try to force more Cooper pairs through, it just doesn’t work. This non-linear effect is extremely important for all our circuits. If we didn’t have that effect, then we wouldn’t be able to control or manipulate any quantum information that we store in these circuits,” Hays explains.
But sometimes, Cooper pairs can unexpectedly squeeze through the barrier two at a time, an effect that is known as a second-order harmonic correction. This effect limits the performance of a quantum circuit that has been configured to only allow single-pair tunneling.
“If two Cooper pairs tunnel at the same time, then the assumption we used to build our circuit doesn’t apply anymore. We need to fix the circuit so it can handle that,” Kim says.
But before they can fix the circuit, scientists need to know the source and strength of these distortions.
To obtain this information, the MIT researchers fabricated a quantum circuit so it would be very sensitive to these effects. Essentially, the device is designed to suppress the quantum tunneling process of single Cooper pairs, while allowing the two-pair tunneling process to continue.
In this way, they can detect the presence of second-order harmonic corrections and precisely measure their strength.
Straight to the source
They can also use this circuit to pinpoint the source of these harmonics, which helps researchers identify the best way to correct for them.
There are two potential sources of second-order harmonics — one source is intrinsic to the dynamics of the Josephson junction and the other is caused by the wires connecting the junction to other circuit elements.
While prior research had indicated the second-order harmonics could be due to the dynamics of the junction, the MIT researchers found that additional inductance — the tendency to oppose changes in the flow of electric current —from wires in the circuit was the actual source in their devices.
“This is important because, if we know where the second-order harmonic correction is coming from, we can predict how strong it is likely to be, and use that information to engineer more predictable circuits that will hopefully perform better,” Hays says.
In the future, the researchers want to design experiments that more accurately predict how a device will perform when second-order harmonic corrections occur. They also want to study other sources of second-order harmonic corrections and whether those sources could have negative impacts on a circuit under different fabrication conditions.
This work is funded, in part, by the U.S. Department of Energy, the U.S. Co-design Center for Quantum Advantage, the U.S. Air Force, the Korea Foundation for Advanced Studies, and the Intelligence Community Postdoctoral Research Fellowship Program at MIT.
MIT School of Engineering faculty receive awards in winter 2026Faculty members and researchers were honored in recognition of their scholarship, service, and overall excellence.Each year, faculty and researchers across the MIT School of Engineering are recognized with prestigious awards for their contributions to research, technology, society, and education. To celebrate these achievements, the school periodically highlights select honors received by members of its departments, institutes, labs, and centers. The following individuals were recognized in winter 2026:
Arup K. Chakraborty, the John M. Deutch (1961) Institute Professor in the departments of Chemical Engineering, Chemistry, and Physics, and the founding director of the Institute for Medical Engineering and Science, as well as James J. Collins, the Termeer Professor of Medical Engineering and Science in the Department of Biological Engineering and IMES, received the 2026 Laureate of the Tel Aviv University International Prize in Biophysics. The prize recognizes outstanding scientists whose work has significantly advanced the understanding of biological systems through physical principles.
Anantha Chandrakasan, MIT provost and the Vannevar Bush Professor in the Department of Electrical Engineering and Computer Science, received the 2025 IEEE Journal of Solid-State Circuits Test of Time Award. The award recognizes an outstanding paper published in the IEEE Journal of Solid-State Circuits at least 10 years prior that has had significant impact on its field.
Charles Harvey, a professor in the Department of Civil and Environmental Engineering; Piotr Indyk, the Thomas D. and Virginia W. Cabot Professor in the Department of Electrical Engineering and Computer Science; John Henry Lienhard, the Abdul Latif Jameel Professor of Water and Mechanical Engineering in the Department of Mechanical Engineering; Frances Ross, the TDK Professor in Materials Science and Engineering; Zoltán Sandor Spakovszky, the T. Wilson (1953) Professor in Aeronautics; and Ram Sasisekharan, the Alfred H. Caspary Professor of Biological Physics and Physics in the Department of Biological Engineering; were elected to the National Academy of Engineering for 2026. One of the highest professional distinctions for engineers, membership in the NAE is given to individuals who have made outstanding contributions to “engineering research, practice, or education,” and to “the pioneering of new and developing fields of technology, making major advancements in traditional fields of engineering, or developing/implementing innovative approaches to engineering education.”
Michael Howland, the Jeffrey Cheah Career Development Professor and assistant professor in the Department of Civil and Environmental Engineering, received a 2026 Faculty Early Career Development (CAREER) Award from the National Science Foundation. The award supports early-career faculty who have the potential to serve as academic role models in research and education and to lead advances in the mission of their department or organization.
Yoon Kim, associate professor in the Department of Electrical Engineering and Computer Science; Anand Natarajan, an associate professor in the Department of Electrical Engineering and Computer Science; and Mengjia Yan, ITT Career Development Professor in Computer Technology and associate professor in the Department of Electrical Engineering and Computer Science, were named 2026 Sloan Research Fellows. Sloan Research Fellowships support fundamental research conducted by early-career scientists, and they are awarded annually to early-career researchers whose creativity, innovation, and research accomplishments make them stand out as the next generation of leaders.
Carlos Portela, the Robert N. Noyce Career Development Professor and associate professor in the Department of Mechanical Engineering, has received a 2026 Young Investigator Award from the Office of Naval Research. The Young Investigator Program seeks to identify and support academic scientists and engineers who are in their first or second full-time tenure-track or tenure-track-equivalent academic appointment, who have received their doctorate or equivalent degree in the past seven years, and who show exceptional promise for doing creative research.
Ellen Roche, the Abby Rockefeller Mauzé Professor and associate department head for research in the Department of Mechanical Engineering and a professor in the Institute for Medical Engineering and Science, received the 2026 Sony Women in Technology Award with Nature. The award recognizes exceptional early- to mid-career women researchers in technology who through their research are driving a positive impact on society and the planet.
Tess Smidt, an associate professor in the Department of EECS, was named co–principal investigator on a National Science Foundation (NSF) AI Research Institute award and also received a 2025 Department of Energy Office of Science Early Career Research Program Award. The NSF AI Materials Institute (AI-MI) aims to propel foundational AI research past the limitations of existing AI algorithms by pursuing materials discovery and conquering knowledge- and data-centric challenges. The DoE Early Career Research Program provides five-year awards to exceptional early career researchers at U.S. academic institutions, DoE National Laboratories, and Office of Science User Facilities to stimulate new research directions in mission critical areas supported by DoE’s Office of Science.
Antonio Torralba, the Delta Electronics Professor and faculty head of AI+D in the Department of EECS, was elected to the 2025 cohort of Association for Computing Machinery Fellows. ACM Fellows, the highest honor bestowed by the professional organization, are registered members of the society selected by their peers for outstanding accomplishments in computing and information technology and/or outstanding service to ACM and the larger computing community.
Harry L. Tuller, a professor in the Department of Materials Science and Engineering, received The Senior Scientist Award from the International Society for Solid State Ionics. The Senior Scientist Award, the most prestigious award of the International Society for Solid State Ionics, is presented to a senior solid-state ionics researcher who has made outstanding contributions to the science and engineering of solid-state ionics.
Vinod Vaikuntanathan, the Ford Foundation Professor of Engineering in the Department of Electrical Engineering and Computer Science, was named a 2026 fellow of the International Association for Cryptologic Research. ACR has established the IACR Fellows Program to recognize outstanding IACR members for technical and professional contributions.
Rethinking how our brains use categories to make sense of the worldResearchers propose a challenge to the traditional view of how the brain uses its ability to categorize.In the new review article, “Categorization is Baked into the Brain,” cognitive scientists Earl K. Miller, Picower Professor of Neuroscience at MIT, and Lisa Feldman Barrett, university distinguished professor at Northeastern University, contend that categorization is part of a predictive process the brain uses to efficiently meet the body’s needs in a fast-paced, otherwise overwhelming sensory world. In that sense, their paper in Nature Reviews Neuroscience challenges decades of dogma about how and why the brain boils down what it sees, hears, smells, tastes, and feels.
Categories are groups of things that are similar enough to be considered functionally equivalent. When you walk through a neighborhood, you’ll naturally experience the furry, four-legged, barking animal ahead of you as a “dog.” In the classic view of cognition, your brain arrives at that categorization by soaking in lots of basic sensory features of the hound — its shape, its size, the sounds it makes, its behavior — and compares that to some prototype “dog” stored in your memory. Hundreds of milliseconds after the first sensory inputs, you can then decide what you might want to do about the dog.
Barrett and Miller argue that that’s wrong. Instead, they propose that your brain comes prepared for sensory patterns with predictions of the motor action plans that are most likely to achieve the needs and goals you bring to the moment. Those prediction signals can be described as a momentary category that the brain constructs to shape the processing of sensory signals.
From the very start, incoming sensory signals are compressed and abstracted into that category to efficiently select the best predicted plan. If you are in an unfamiliar neighborhood your brain might construct the category “dog” to avoid being bit, resulting in: “Back away slowly while saying nice doggie.” If you are on your own block and encounter a familiar dog, your brain might construct a category to kneel and open up your arms to summon your neighbor’s adorable pup for a satisfying petting.
In either case, the category “dog” arises in the context of your needs and your prediction from a menu of learned action plans for similar situations, not from an intellectual exercise of neutrally regarding sensory inputs, comparing them to a fixed prototype, and then planning from there. If the brain really worked the classically believed way, you’d be on the back foot when the unfamiliar dog lunged at you.
“One of the main things your brain has to do is predict the world,” says Miller, a faculty member of The Picower Institute for Learning and Memory and the Department of Brain and Cognitive Sciences at MIT. “It takes several hundred milliseconds to process things, and meanwhile the world is moving on. Your brain has to anticipate things.”
The most pragmatic and efficient way to survive and thrive in such a world, Barrett says, is to have your needs and potential plans ready for the sensory situation. If your predictions are right, you’re prepared in time. If they are wrong, you adjust and learn from it.
“The stimulus, cognition, response model of the brain is wrong,” says Barrett, a faculty member in Northeastern’s Department of Psychology and co-director of the Interdisciplinary Affective Science Laboratory. “The brain prepares for a response and then perceives a stimulus. A brain is not reactive. It’s predictive. Action planning comes first. Perception comes second, as a function of the action plan.”
Anatomical and functional evidence
Throughout the review, Barrett and Miller ground the provocative proposal in copious anatomical, electrophysiological, and imaging evidence about the brain. They cite numerous experimental results that show how the brain is structured to broadcast memories to create motor plans that flow back toward signals that arrive from the body’s sensory surfaces, actively whittling them down and shaping them to give them meaning.
“The capacity to create similarities from differences — to abstract — is embedded in the architecture of the nervous system, and you can see that by looking at what is connected to what and by observing signal flow,” Barrett says.
For example, as circuits feed signals “forward” from sensory surfaces (such as the retina) to regions of the cerebral cortex that are focused on sensory processing (such as the visual cortex) toward the areas that are important for executive control (the prefrontal cortex) and control of the body (limbic cortex), information passes from many small, barely connected neurons to fewer, bigger, and more well-connected neurons. Such an architecture compresses sensory details into increasingly abstract representations that group many different features into smaller groups of similar features, and in doing so helps to select a predicted action plan from the broader category that’s already there.
“Your brain is a big funnel to take the outside world and turn it into an output,” Miller says.
Moreover, anatomical evidence shows that the neurons in the cortex maintain many more connections to provide feedback from memory that control sensory regions than to feed sensory information forward. As much as 90 percent of synapses in the visual cortex are “feedback” instead of “feedforward,” Barrett and Miller wrote. In other words, the brain is built to use memory to filter incoming sensory signals, consistent with imposing needs and goals on what would otherwise be a deluge of sights, sounds, and other sensations.
Yet another line of evidence are numerous studies from Miller’s own lab showing that at the broad network level of information flow in the cortex, the brain uses beta frequency waves that carry information about goals and plans, to constrain the expression of gamma frequency waves that carry information about specific sensory inputs.
Finally, the dominance of “feedback” over “feedforward” signals in the cortical architecture allows for the possibility that sensory signals are made meaningful in terms of predicted plans. When these plans are wrong, the resulting surprise can be integrated for future use.
“In science, there is a special name for that: learning,” Barrett says.
Implications for human thought and disease
In the end, Barrett and Miller’s proposal completely changes the idea of categorization, shifting it from being a particular intellectual skill to being a fundamental function for predictively meeting the body’s needs (or, “allostasis”).
“A category may not be a representation that an animal has, but a signal processing event than an animal does, predictively, to constrain the meaning of a high-dimensional ensemble of signals in a particular situation,” the authors wrote. “Categorization renders these signals meaningful — similar to one another and to past allostatic events — in terms of some goal or function.”
Humans, Barrett says, have a relatively massive amount of the neural network architecture to perform these pragmatic abstractions, and therefore can make categorizations that seem outright metaphorical (e.g., a functional similarity between “climbing the career ladder” and climbing a literal physical ladder).
But these processes can also go awry in disease, Barrett and Miller note. Depression can be seen as a disorder in which the brain imposes overly broad categories, such as “threat” or “criticism” on sensory episodes that don’t have to be perceived that way. By contrast, autism can manifest with features of inadequate compression of incoming sensory signals, not generalizing enough to recognize when a situation is similar enough to a prior one to select the appropriate plan.
Funding to support the paper came from the National Institutes of Health, The U.S. Army Research Institute for the Behavioral and Social Sciences, the Office of Naval Research, the Unlikely Collaborators Foundation, The Freedom Together Foundation, and The Picower Institute for Learning and Memory.
MIT BrainTrust supports neighbors living with brain injuriesNearly 100 MIT students participate in a buddy program that assists Boston-area residents.Since 1998, members of MIT’s BrainTrust club have helped Boston-area residents with brain injuries or other neurological disorders through their buddy program. The organization’s members also visit patients in nursing homes suffering from neurological issues.
BrainTrust is one of the founding chapters of Synapse National, an organization created by MIT alumna Alissa Totman ’13. Synapse’s goal is to provide social support for individuals living with brain injuries and to educate and inspire student leaders in the field of brain injury.
“Learning directly from individuals who had experienced brain injury during my time in BrainTrust gave me an appreciation of the gaps in resources and opportunities for improvement in brain injury care, which ultimately motivated me to pursue a career in brain injury medicine. My experience in BrainTrust continues to shape my approach to patient care and my professional goal of improving access to specialized care for individuals with brain injury by serving as a consulting provider in the acute care hospital, as well as by training the next generation of leaders in the field,” says Totman.
The club’s president, junior Karie Shen, who is pursuing a double major in biology (Course 7) and brain and cognitive science (Course 9), says, “BrainTrust is a student-run service organization that provides support for individuals with brain injury and other neurological disorders. I joined BrainTrust because it seemed like the perfect intersection of community service and neuroscience, and I care about these two things deeply.”
BrainTrust volunteers participate in training and then are paired with a local buddy who has experienced a brain injury. Members can also spend time on the weekends with patients in nursing homes who have dementia, Alzheimer’s disease, or who have had a stroke.
Shen, along with Elizabeth Zhang, president of the MIT Pre-Med Society, recently developed a program that allows BrainTrust members to visit patients in hospice. “It’s an experience that is deeply valuable for students. We work through a third-party organization called Compassus. Because the pairing process is HIPAA-protected, our role as BrainTrust executive members is to recruit students and connect them with the hospice volunteer coordinator for training. We also provide funding for transportation, generously supported by the UA Community Service Committee,” says Shen.
Shen, who plans to go to medical school and specialize in neurology, neuro-oncology, or geriatric medicine when she completes her degree, finds the experience rewarding, at times difficult, but also offers a glimpse into the reality of working with people with brain injuries.
“Visiting the people in hospice or a nursing home is hard. I’ve seen residents cry for no apparent reason that the nurses or I can understand. But I have also come to understand that caring for a patient’s quality of life and dignity is equally important. What I came to realize is that my presence itself mattered. That perspective has shaped how I think about the kind of physician I want to become,” says Shen.
First-year student Jordan Lacsamana heard about the club during Campus Preview Weekend and was immediately interested. Lacsamana, who will major in brain and cognitive sciences, is a volunteer in the Buddy Program and meets with her buddy at least once a month.
“I joined the club because it aligned with my interests academically, but I also wanted to support someone in the Boston community. I’m pre-med, and I’m interested in surgery, possibly neurosurgery or cardiovascular surgery. But I also think it’s nice to have someone outside of MIT to talk with. It’s great to learn more about them and have that one-on-one friendship, which really is the goal,” says Lacsamana.
Lacsamana says she enjoys spending time with Amanda, her buddy, and exploring Boston and Harvard Square, meeting for coffee or meals, and getting as much out of the relationship as Amanda does.
“I see her as a mentor because coming to Boston from Dallas was such a big change, so I’ve also been able to look to her for advice. But I think one of the great things about the program is that you get to learn more about them as an individual, instead of seeing them as just a person with an injury,” says Lacsamana.
“Many of our brain injury buddies simply enjoy being around students, staying connected to what we are learning and doing. Some have been with the club for years, even upwards of a decade, and still keep up with former student members long after they graduate. It is really wonderful to see how BrainTrust has created this web of friendships between people who would otherwise never have met,” says Shen.
“Amanda has stayed in touch with her former buddy since she graduated from MIT and is going to her wedding,” says Lacsamana. “I think it’s a testament to how amazing this program is at forming those connections.”
MIT students who seek real-world opportunities in fields such as cognitive science, health care, medicine, and cognitive/neurological prosthetics, or who want to help a local resident, can join BrainTrust. Email braintrust-exec@mit.edu for more information.
Astronomers pin down the origins of a planetary odd coupleNew measurements of a hot Jupiter and its mini-Neptune companion suggest both planets formed surprisingly far away from their host star.Across the Milky Way galaxy, a planetary odd couple is circling a star some 190 light years from Earth. A normally “lonely” hot Jupiter is sharing space with a mini-Neptune, in a rare and unlikely pairing that’s had astronomers puzzled since the system’s discovery in 2020.
Now MIT scientists have caught a glimpse into the atmosphere of the mini-Neptune, which is circling inside the orbit of its Jupiter-sized companion, and discovered clues to explain the origins of this unusual planetary system.
In a study appearing today in Astrophysical Journal Letters, the scientists report on new measurements of the mini-Neptune’s atmosphere, made using NASA’s James Webb Space Telescope (JWST). It is the first time astronomers have measured the composition of a mini-Neptune that resides inside the orbit of a hot Jupiter.
Their measurements reveal that the smaller planet has a “heavy” atmosphere that is rich with water vapor, carbon dioxide, sulfur dioxide, and hints of methane. Such a heavy atmosphere would not have been acquired by the planet if it had formed in its current location, very close to its star.
Instead, the scientists say their findings point to an alternate origin story: Both the mini-Neptune and the hot Jupiter may have formed much farther away, in the colder region of the protoplanetary disk. There, the planets could slowly build up atmospheres of ice and other volatiles. Over time, the planets were likely drawn in toward the star in a gradual process that kept them close, with their atmospheres intact.
The team’s results are the first to show that mini-Neptunes can form beyond a star’s “frost line.” This boundary refers to the minimum distance from a star where the temperature is low enough that water instantly condenses into ice.
“This is the first time we’ve observed the atmosphere of a planet that is inside the orbit of a hot Jupiter,” says Saugata Barat, a postdoc in MIT’s Kavli Institute for Astrophysics and Space Research and the lead author of the study. “This measurement tells us this mini-Neptune indeed formed beyond the frost line, giving confirmation that this formation channel does exist.”
The team consists of astronomers around the world, including Andrew Vanderburg, a visiting assistant professor at MIT, and co-authors from multiple other institutions including the Harvard and Smithsonian Center for Astrophysics, the University of South Queensland, the University of Texas at Austin, and Lund University.
A “one-of-a-kind” system
As their name implies, mini-Neptunes are planets that are less massive than Neptune. They are considered to be gas dwarfs, which are made mostly of gas, with an inner, rocky core. Mini-Neptunes are the most commonly found planet in the Milky Way, though, interestingly, no such world exists in our own solar system. Astronomers have observed many planets circling a wide variety of stars in a range of planetary systems. Mini-Neptunes, then, are generally considered to be garden-variety planets.
But in 2020, Chelsea X. Huang, then a Torres Postdoctoral fellow at MIT (now on the faculty at University of South Queensland), discovered a mini-Neptune in a rare and puzzling circumstance: The planet appeared to be circling its star with an unlikely companion — a hot Jupiter.
The astronomers made their discovery using NASA’s Transiting Exoplanet Survey Satellite (TESS). They analyzed TESS’ measurements of TOI-1130, a star located 190 light years from Earth, and detected signs of a mini-Neptune and a hot Jupiter, orbiting the star every four and eight days respectively.
“This was a one-of-a-kind system,” says Huang. “Hot Jupiters are ‘lonely,’ meaning they don’t have companion planets inside their orbits. They are so massive, and their gravity is so strong, that whatever is inside their orbit just gets scattered away. But somehow, with this hot Jupiter, an inner companion has survived. And that raises questions about how such a system could form.”
A spot-on snapshot
The 2020 discovery of TOI-1130 and its odd planetary pair inspired Huang, Vanderburg, and their colleagues to take a closer look at the planets, and specifically, their atmospheres, with JWST. In its new study, the team reports its analysis of TOI-1130b — the inner-orbiting mini-Neptune.
Catching the planet at just the right time was their first challenge. Most planets circle their star with a regular, predictable period, like the tick of a clock. But the mini-Neptune and the hot Jupiter were found to be in “mean motion resonance,” meaning that each can affect the other’s motion, pulling and tugging, and slightly varying the time each takes to orbit their star. This made it tricky to predict when JWST could get a clear view.
The team, led by Judith Korth of Lund University, assembled as many past observations of the system as they could, and developed a model to predict when each planet would pass by the star at an angle that JWST could observe.
“It was a challenging prediction, and we had to be spot-on,” Barat says.
In the end, the team was able to catch a direct and detailed snapshot of both planets.
“The beauty of JWST is that it does not observe just in one color, but at different colors, or wavelengths,” Barat explains. “And the specific wavelengths that a planet absorbs can tell you a lot about the composition of its atmosphere.”
From JWST’s measurements, the team found that the planet absorbed wavelengths specifically for water, carbon dioxide, sulfur dioxide, and to a lesser degree, methane. These molecules are heavier than hydrogen and helium, which constitute lighter atmospheres. Astronomers had assumed that, if mini-Neptunes formed very close to their star, they should have light atmospheres.
But the team’s new results counter that assumption and offer a new way that mini-Neptunes could form. Since heavier molecules were found in the atmosphere of TOI-1130b, which resides very close to its star, the scientists say the only possible explanation for its composition is that the planet formed much farther out than its current location.
The planet likely accumulated its heavy atmosphere of water and other volatiles such as carbon dioxide and sulfur dioxide in the icy region beyond the star’s frost line. In this much colder environment, water condenses onto bits of dust to form icy pebbles, which an infant planet can draw into its atmosphere. The water evaporates as it slowly migrates in closer to its star.
Barat says the team’s detection of heavy molecules in the atmosphere of TOI-1130b confirms that the planet — and likely its hot Jupiter companion — formed in the outskirts of the system. Through gradual migration, the two planets would be able to stay close together and keep their atmospheres intact.
“This system represents one of the rarest architectures that astronomers have ever found,” Barat says. “The observations of TOI-1130b provide the first hint that such mini-Neptunes that form beyond the water/ice line are indeed present in nature.”
This work was supported, in part, by NASA.
Biologist Joey Davis explores how cells build complex structures His studies have shed light on the assembly instructions that govern ribosomes, the critical protein-building machines of the cell.Ribosomes, the cellular machines that assemble proteins, are made from dozens of proteins and RNA molecules. Putting all of those pieces together is a complex puzzle — one that MIT Associate Professor Joey Davis PhD ’10 revels in trying to solve.
Understanding how these structures form and later break down could help researchers learn more about how disruptions of these fundamental processes can lead to disease. But, as Davis points out, it’s also an interesting biological question.
“Our long-term goal is to really understand how the natural world assembles these huge complexes rapidly and efficiently. It’s a fundamentally interesting question to think about how these things get put together,” he says.
His work has helped reveal that unlike building a house, which happens in a prescribed sequence of steps — pouring the foundation, building the frame, putting on the roof, then doing electrical and plumbing work — ribosomes can be assembled in a more flexible way. Cells can even skip an assembly step and then come back to it later.
“In these natural systems, it seems like the assembly pathways are much more dynamic and flexible,” he says. “It appears that evolution has selected pathways that aren’t strictly ordered in the way we would think about an assembly line, where you always put in one component, then the next, and then the next. We’re excited to understand the selective advantages of such approaches.”
A love of discovery
Davis’ interest in how things are put together developed early in life, inspired by his father, a carpenter who framed houses. During the mid-1980s, the family moved from Colorado to Southern California, where his father worked in construction during a housing boom there.
“I was always interested in building things, which I think probably came from being around my dad and other builders,” Davis says.
As an undergraduate at the University of California at Berkeley, where he majored in computer science and biological engineering, Davis’ interests turned toward smaller scales, in the realm of cells and molecules. During his junior year, he started working in the lab of chemistry professor Michael Marletta, who studies molecular-level biological interactions.
In the lab, Davis investigated how enzymes that contain heme are able to preferentially bind to either oxygen or nitric oxide, two gases that are very similar in structure. That work kindled a love of studying the natural world and pursuing discoveries in fundamental science.
“Being in the Marletta lab and seeing students and postdocs that were really passionate about these problems had a big impact on me,” Davis says. “The goal was to understand the fundamentals of how molecular discrimination works, and the idea of discovery for the sake of discovery was thrilling.”
After graduating from Berkeley, Davis spent another year working in Marletta’s lab, and then a year working odd jobs, before heading to MIT to pursue a PhD in biology. There, he worked with Professor Bob Sauer, now emeritus, who studied the relationship between protein structure and function, with a particular focus on the molecular machines that degrade or remodel proteins.
Davis’ thesis research centered on enzymes called AAA proteases, which remove damaged proteins from cellular membranes and send them to cell organelles that break them down. In addition to studying the structure and function of the proteases, Davis worked on ways to engineer them to tag specific proteins for destruction.
That work led him into synthetic biology, which he used to develop genetic parts that drive production of proteins of interest. Some of those parts ended up being used by the biotech startup Ginkgo Bioworks, where Davis took a job as a senior scientist after graduating.
Working at Ginkgo Bioworks allowed Davis to stay in Boston while his partner finished her PhD. The couple then moved back to California, where Davis worked as a postdoc at Scripps Research, which was home to one of the first direct electron detection cameras for cryo-electron microscopy (cryo-EM). These detectors allow researchers to generate structures with near atomic resolution. At Scripps, Davis began using them to study ribosomes as they were being assembled.
Peering into the ribosome
After joining the MIT faculty in 2017, Davis continued his work on ribosomes and assembled a lab group that includes students from a variety of backgrounds who work together to develop new ways to explore biological phenomena.
“I have a mix of method developers and biologists in the group, and the work from each of them informs each other,” Davis says. “My lab goes back and forth between building sets of tools to answer biological questions, and then as we’re answering those questions, it motivates the next generation of tool development.”
During ribosome assembly, RNA molecules fold themselves into the correct shapes, creating docking sites for proteins to attach. Then, more RNA molecules come in and fold themselves into the structure.
“It’s a beautifully coupled process by which the cell folds hundreds of RNA helices and binds on the order of 50 proteins, and it does it in two minutes from start to finish. E. coli does this 100,000 times per hour, and it’s amazing how rapid and efficient the process is,” Davis says.
Cryo-EM allows scientists to capture this process in minute detail. It can be used to take hundreds of thousands of two-dimensional images of ribosome samples frozen in a thin layer of ice, from different angles. Computer algorithms then piece together these images into a three-dimensional representation of the ribosome.
To gain insight into how ribosomes are assembled, researchers can stall the process at different points and then analyze the resulting structures. In 2021, Davis’s lab developed a new method called CryoDRGN, which uses neural networks to analyze cryo-EM data and generate the full ensemble of structures that were present in the sample.
This work has shown that when certain steps of ribosome assembly are blocked, many different structures result, suggesting that the assembly can occur in a variety of ways.
In future work, Davis aims to dramatically increase the throughput of cryo-EM to generate datasets of protein structures that could help improve the AI-based models that are now used to predict protein structures.
“There are still huge swaths of sequence space that these models are very poor at predicting, but if we could collect data on those sequences en masse, that could potentially serve as key training data for a next-generation protein structure prediction method that could fill out that space,” he says.
Rett syndrome study highlights potential for personalized treatmentsUsing advanced human cell cultures, MIT researchers tracked how two different mutations alter neural circuit development, and how each could be addressed with distinct potential therapeutics.Although many studies approach the developmental disorder Rett syndrome as a single condition arising from general loss of function in the gene MECP2, a new study by neuroscientists in The Picower Institute for Learning and Memory at MIT shows that two different mutations of the gene caused many distinct abnormalities in lab cultures. Moreover, correcting key differences made by each mutation required different treatments.
“Individual mutations matter,” says Mriganka Sur, senior author of the new open-accdess study in Nature Communications and the Newton Professor in the Picower Institute and the Department of Brain and Cognitive Sciences. “This is an approach to personalizing treatment, even for a single-gene disorder.”
The study employed advanced 3D human brain tissue cultures called “organoids” or “minibrains” derived from skin cells or blood cells donated by Rett syndrome patients with each mutation. Lead author Tatsuya Osaki, a Picower Institute research scientist, says that the organoids’ ability to model the specific consequences of each mutation enabled him to gain mutation-specific insights that haven’t emerged in prior studies, where scientists just knocked out MECP2 overall. The organoids also provided a novel opportunity to understand how each mutation affected different cell types and their interactions.
Distinct effects
More than 800 mutations in MECP2 can cause Rett syndrome, but just eight account for more than 60 percent of cases. Sur and Osaki chose one of these, R306C, which involves a difference of just one DNA base pair (916C>T), because it represents 7-8 percent of Rett syndrome cases. The other mutation they chose, V247X, is much more rare and severe because it cuts off production of the gene’s protein product by a single DNA base deletion (705Gdel), leaving the protein not just errant, but incomplete.
In organoids cultured for three months, each mutation produced some common but also sometimes distinct consequences compared to control organoids with non-mutated MECP2. For many of their experiments, the team used “three-photon” microscopes capable of cellular-level resolution all the way through the organoids’ approximate 1 millimeter thickness, resolving both their structure (via “third-harmonic generation” imaging), and the live activity patterns of their neurons (via calcium fluorescence).
For instance, the scientists observed that the V247X organoids exhibited several structural differences from their controls — they were larger and had different thicknesses of various layers — but the R306C ones were much more like their controls. Organoids harboring either mutation exhibited less-developed axon projections from their neurons, compared to their control comparators.
Looking at properties of neural activity and connectivity in the organoids, the scientists found some similar deficits across both mutations. Both showed reduced spiking activity and synchronicity between neurons compared to in their controls.
But when the scientists looked at other properties, the organoids started to diverge from each other. In particular, an indication of the efficiency of their network structure called “small-world propensity” (SWP) was decreased in R306C organoids, and increased in V247X ones, compared to controls. This means that both mutations altered the development of typical network structures for information processing, but in different directions.
To ensure that their results were meaningful for Rett syndrome patients, the team collaborated with Charles Nelson at Boston Children’s Hospital, whose team measured EEG in several children with different Rett mutations. Although the sample was small, the researchers measured indications that the SWP property in the EEG readings was altered in the volunteers, much like in the organoids.
Finally, by labeling excitatory neurons to flash in one color and inhibitory neurons to flash in a different color, the scientists were able to see that connectivity between the different neural types differed significantly from controls in the V247X organoids.
Treatment tests
All the testing showed that each mutation caused several changes in organoid structure, activity, and connectivity, and that the deviations were often particular to the specific mutation.
To understand how these differences emerged, and how they might be corrected, Sur and Osaki’s team turned to examining how the cells in each kind of organoid might be expressing their genes differently than controls. Differences in gene expression often lead to alterations of key molecular pathways in cells that can disrupt their activity and function. Analysis with a technique called single cell RNA sequencing indeed yielded hundreds of differences in each organoid type, where some genes were expressed more than in controls while others were underexpressed.
For instance, the analyses revealed that in R306C organoids a gene called HDAC2 was overexpressed. That protein is known for repressing expression of other genes. Meanwhile, in the V247X organoids, the scientists found reduced expression of genes for some receptors of the inhibitory neurotransmitter GABA. These organoids also showed defects in the function of astrocyte cells, which support many aspects of neural function.
Organoids with either mutation also exhibited aberrations in molecular pathways that enable the development of circuit connections between neurons, called synapses.
Given the specific defects they observed, the scientists decided to treat the organoids with a drug that can inhibit HDAC2 activity and another that increases GABA’s efficacy. The HDAC2 inhibitor restored neuronal activity and SWP to normal levels in the R306C organoids, and the GABA “agonist” baclofen restored SWP to control levels in the V247X organoids.
Osaki notes each of the treatment drugs has already been studied in other disease contexts, meaning they are well-understood drugs that could be repurposed.
Now that the researchers have developed an organoid platform for dissecting individual mutations’ consequences, identifying both their roots and testing treatments, they plan to apply it to studying four more mutations, Sur says, comparing all of them against a standardized control organoid.
In addition to Sur, Osaki, and Nelson, the paper’s other authors are Chloe Delepine, Yuma Osako, Devorah Kranz, April Levin, and Michela Fagiolini.
The National Institutes of Health, a MURI grant, The Freedom Together Foundation, and the Simons Foundation provided support for the research.
Powering 160,000 hours of discovery at MIT.nanoNanoFab Equipment Management and Operations (NEMO) system streamlines shared facilities management via tool trainings, reservations, and lab communications.Each year, more than 1,500 researchers rely on over 200 tools and instruments at MIT.nano to pursue experiments that span MIT’s disciplines, collectively generating 160,000 hours of work across 88,000 instances of tool use. Behind this activity is an operational framework that must discretely coordinate access, maintain fairness, and keep research moving without friction.
Managing such a dynamic environment requires more than a scheduling calendar. An automated reservation system serves as the connective tissue of the facility, balancing demand across diverse user needs while supporting the practical realities of a shared lab space. Researchers arrive at MIT.nano with different workflows, safety requirements, and administrative needs, yet the system must present a seamless experience. Integration with MIT’s broader digital infrastructure, from onboarding and authentication to safety training and billing, ensures that access is both efficient and compliant, reducing barriers so researchers can focus on their work.
A system for the modern era
Over the past three years, during a period of rapid growth in both equipment and facility usage, MIT.nano undertook a transition to a new platform designed to scale with demand while maintaining operational continuity. The effort reflects an ongoing commitment to evolving infrastructure that supports the pace, complexity, and collaborative spirit of modern research.
The importance of robust laboratory management systems has long been recognized at MIT. For decades, researchers in the Microsystems Technology Laboratories (MTL) and the Materials Research Laboratory relied on the CORAL lab management platform to reserve and manage shared instrumentation. Jointly developed by MIT and Stanford University and introduced in 2003, CORAL represented a significant advance over the text-based system it replaced. But by the time MIT.nano adopted CORAL in 2018, active development had slowed, and the platform was beginning to show its age, most visibly through the absence of modern web and mobile interfaces expected by today’s users.
To address these limitations, MIT.nano has transitioned to NEMO, an open-source laboratory management system originally developed at the National Institute of Standards and Technology. NEMO centralizes scheduling, communication, and operational logistics into a single platform that manages tool reservations and user access while supporting facility growth. Its modular architecture and plugin framework allow for extensive customization, enabling the system to evolve alongside the needs of a large, shared research environment.
“Over time, NEMO was replicating core functionalities of CORAL while introducing new features that CORAL simply could not support,” explains Thomas Lohman, senior software and systems manager at MTL and a long-time contributor to CORAL’s development. “The question became whether to continue patching the old system or adopt this new platform that already had a lot of the features we use daily, as well as an active community continually improving it.”
For MIT.nano leadership, modernization was about more than replacing an aging tool. “We needed a system that centralizes everything a facility user depends on — policies, tool documentation, training workflows, and communications — within a user-friendly, mobile-accessible environment,” says Anna Osherov, associate director for Characterization.nano, who led the evaluation and transition effort. “Just as important was making sure the platform enhances the experience for both users and staff.”
Collaborating at MIT and with shared access facilities
MIT.nano collaborated closely with Mathieu Rampant, NEMO project lead and CEO of Atlantis Labs, to adopt the community edition of NEMO, an extended version enriched by contributions from a growing global user base. The open-source model ensures that improvements developed at MIT.nano benefit the broader research community, reinforcing a shared ecosystem of innovation. “The NEMO community is expanding rapidly, and many new features originate directly from facility users and administrators,” says Rampant. “That collaborative model allows improvements to propagate quickly while giving institutions a sense of ownership in the platform’s evolution.”
NEMO introduces modern features long requested by MIT.nano researchers, including mobile access, improved transparency, and streamlined workflows. Facility users can now monitor their own tool usage and consumables, customize notifications, register for training, join real-time equipment waitlists, report issues, and communicate with staff, all through a unified dashboard. What was once distributed across multiple systems is now centralized, reducing friction in day-to-day lab operations.
Launching a new platform at the scale of MIT.nano required careful planning and sustained collaboration. The system needed to support multiple facility types, integrate with existing MIT infrastructure, and accommodate a diverse set of instrumentation workflows. “Features that work well in a typical characterization lab can quickly become a burden in a more chemically active environment like the cleanroom,” explains Jorg Scholvin, associate director of Fab.nano. “Relying on researchers to log in using personal devices and Duo authentication, for example, would be impractical in that setting.”
To address these challenges, MIT.nano collaborated with MIT Information Systems and Technology Associate Vice President Olu Brown and Senior Director for Infrastructure Operations Marco Gomes and their teams to streamline integration between MIT systems and NEMO for cleanroom users. “The availability of modern APIs allowed us to connect very different systems efficiently and deliver a convenient, seamless, and productive experience in the lab,” says Scholvin.
The result is a platform that now processes thousands of reservations, communications, and operational actions daily. “We truly value the partnership with MIT.nano and appreciate the collaboration throughout this effort,” says Gomes. “It’s been a great example of teams working together to deliver something meaningful for the research community.”
As one of the largest shared-access facilities deploying NEMO, MIT.nano has played a central role in advancing the platform’s capabilities, both by helping shape its development and by demonstrating a model that is scalable and effective for other facilities and research centers nationwide. Enhancements first created to meet MIT.nano’s needs are now leveraged by other facilities adopting NEMO across the globe.
Beacon Biosignals is mapping the brain during sleepFounded by Jake Donoghue PhD ’19 and former MIT researcher Jarrett Revels, the company is creating an AI-driven platform to help diagnose and treat disease.The human brain remains one of the most fascinating and perplexing mysteries in medicine. Scientists still struggle to match neurological activity with brain function and detect problems early, slowing efforts to treat neurological disorders and other diseases.
Beacon Biosignals is working to make sense of the brain by monitoring its activity while people sleep. The company, which was founded by Jake Donoghue PhD ’19 and former MIT researcher Jarrett Revels, developed a lightweight headband that uses electroencephalogram (EEG) technology to measure brain activity while people enjoy their normal sleep routines at home. Those data are processed by machine-learning algorithms to monitor the effects of novel treatments, find new signs of disease progression, and create patient cohorts for clinical trials.
“There’s a step-change in what becomes possible when you remove the sleep lab and bring clinical-grade EEG into the home,” says Donoghue, who serves as Beacon’s CEO. “It turns sleep from a constrained, facility-based test into a scalable source of high-quality data for diagnostics, drug development, and longitudinal brain health.”
Beacon partners with pharmaceutical companies to accelerate its path to patients. The company’s FDA 510(k)-cleared medical device has already been used in over 40 clinical trials across the globe as part of studies aimed at treating conditions including major depressive disorder, schizophrenia, narcolepsy, idiopathic hypersomnia, Alzheimer’s disease, and Parkinson’s disease.
With each deployment, Beacon learns more about how the brain works — insights it is using to create a “foundation model” of the brain.
“It’s our belief that the dataset that’s going to transform brain health doesn’t exist yet — but we are rapidly creating it,” Donoghue says. “Our platform can characterize the heterogeneity of disease progression, generating dynamic insights that are impossible to fully capture through static modalities like sequencing or imaging. The brain is an electric organ and changes through synaptic plasticity, so tracking brain function across many diseases at scale will allow us to discover novel subgroups of diseases and map them over time.”
Illuminating the brain
Donoghue trained in the Harvard-MIT Program in Health Sciences and Technology, conducting clinical training for an MD while completing his PhD in neuroscience at MIT under the guidance of Earl Miller, MIT's Picower Professor in Brain and Cognitive Sciences and The Picower Institute for Learning and Memory. While in the program, Donoghue trained at Massachusetts General Hospital and Boston Children’s Hospital, where he helped care for patients, including in oncology, during the rise of genomic sequencing to guide precision cancer therapies. He later worked in neurology and psychiatry, where care often relied on more iterative approaches — highlighting an opportunity to bring similarly data-driven precision to brain health.
“What struck me most was the inability to measure brain function in the ways that cardiologists can longitudinally monitor cardiac function in patients from home,” Donoghue says. “At MIT, I built this conviction that processing a lot of brain data and working to correlate that with brain function would be transformative to how these neurological diseases are identified and treated.”
Toward the end of his training, Donoghue began developing his ideas further, engaging with mentors including HST and Harvard Medical School professors Sydney Cash and Brandon Westover. He had met Revels, who was working as a research software engineer in MIT’s Julia Lab, during his PhD, and convinced him to co-found Beacon with him in 2019.
“We decided building a business to understand the organ of interest — the brain — would be a great start to understanding heterogeneous neuropsychiatric diseases and building better treatments,” Donoghue recalls.
Beacon began as a computation and analytics company building wearable devices to expand clinical impact and reach. From its early days, Beacon has been partnering with large pharmaceutical companies running clinical trials, offering a less invasive way to watch brain activity and learn how their drugs are impacting the brain as well as how patients sleep.
“It was clear sleep was the right window to understand the brain,” Donoghue says. “Neural activity during sleep can be an order of magnitude higher and more structured, almost like a language. It’s a great surface area for understanding brain function and how different drugs affect the brain.”
Donoghue says Beacon’s devices can collect lab-grade data on each patient for multiple sequential nights, resulting in higher quality assessment. The company uses machine learning to extract insights, such as the time patients spend in different sleep stages and the number of small awakenings that occur throughout the night. It can also detect subtle sleep architecture changes that might lead to cognitive decline.
“We’re starting to take features of sleep activity and link them to outcomes in a way that’s never been done with this level of precision,” Donoghue says.
To date, Beacon has taken part in clinical trials for sleep and psychiatric disorders as well as neurodegenerative diseases, where sleep changes can emerge years before the presentation of symptoms.
“We do a lot of work in areas like Alzheimer’s disease and Parkinson’s, which affected my grandfather,” Donoghue says. “We’re analyzing features of rapid-eye-movement and slow-wave sleep to detect early changes that precede clinical symptoms. It’s an opportunity to move these diseases from late recognition to much earlier, data-driven detection.”
Improving brain treatments for millions
Last year, Beacon acquired an at-home sleep apnea testing company that serves more than 100,000 patients each year across the U.S., accelerating access to high-quality, comprehensive testing in the home and expanding the reach of its platform. Then in November, the company raised $97 million to accelerate that expansion.
“The vision has always been to reach patients and help people at scale,” Donoghue says. “What’s powerful is that we’re building a longitudinal record of brain function over time,” Donoghue says. “A patient might come in for sleep apnea screening, but if they develop Parkinson’s years later, that earlier data becomes a window into the disease before symptoms emerged. That turns routine testing into a foundation for entirely new prognostic biomarkers — and a path to detecting and intervening in brain disease earlier, potentially before symptoms ever begin.”
Improving understanding with language MIT senior Olivia Honeycutt investigates how the ways we communicate can shape our views of the world.When she was a child, MIT senior Olivia Honeycutt would spend summers on her grandparents’ farm in rural Alabama outside Birmingham. The practical and cultural differences between farm and city life became more pronounced by comparison. “Life and the way we lived it slowed down on the farm,” she says. “It was a nice change of pace.”
These days, Honeycutt, a double major in computation and cognition and linguistics, still finds herself moving between several worlds that are simultaneously connected and distinctly different. Her research interests lie at the intersection of human thinking and awareness, language learning and acquisition, technology, and social group interaction and impact.
Honeycutt’s interest in language and the ways it can shape how we think and live grew alongside lifelong investments in math and science. She learned French from her relationships with Haitian family friends, and American Sign Language because of another friend’s deaf sibling. She was fascinated with how speakers from those groups communicated and how the brain can reorganize itself when confronted with a lack of auditory input.
“There are so many things that are different about sign language and spoken language,” she says. “Speaking in multiple languages and dialects while managing the emotional and cultural nuances multilingualism presents can shift your experience of the world and of yourself.” Operating in these areas creates research opportunities in disciplines as diverse as neurology, large language models (LLMs), psychology, and public policy.
“There’s fascinating work underway in neurolinguistics,” Honeycutt notes, “along with trying to better understand the differences between neural networks, AI, and how each processes information.” She’s wanted to study these for a long time, she says. “When people have to manage language deficits like aphasia, for example, and you’re immersed in several areas of investigation to find answers, you get to learn cool things like how the brain ‘does’ language.”
An MIT approach to study
Honeycutt chose MIT, in part, because the computation and cognition major was “not something I could find elsewhere.” Her affinity for math and English, alongside a desire to pursue the kind of computer science work that “centered people,” increased the likelihood that she could continue in her preferred areas of investigation with the support of the Institute’s faculty and other students.
She found class 9.59J (Laboratory in Psycholinguistics), taught by professor of brain and cognitive sciences Ted Gibson, to be especially enlightening. “It laid the foundation for my work,” she says.
Her decision to major in linguistics along with computation and cognition meant she could connect her interests in brain function and technology with a data-driven approach to language study and processing. “Majoring in linguistics highlighted the power of scientific rigor to organize and analyze a vast amount of chaotic, human-centric data,” she says. Her coursework reinforced the value of her decision.
Honeycutt lauds the freedom MIT’s focus on interdisciplinary study provides. “Researchers are exploring differences between human and LLM language models and processing, and a lot of that work is happening at MIT,” she says. “MIT provides a rigorous flexibility that allows me to indulge multiple academic interests.”
It’s this flexibility that Honeycutt values most. “It’s the only reason I’m on the path I’ve chosen,” she continues, one that features a focus on language acquisition, education policy, LLMs’ computational possibilities and limitations, and education reform.
Honeycutt’s research continued on a series of MISTI trips in 2025. She traveled to South Africa in the summer, where she worked on the South African Human Rights Commission’s “Right to Read” campaign. She explored connections between language processing and brain function and supported research to aid in developing legislation to help increase literacy among South Africans.
“Linguistic diversity presents significant challenges in South Africa,” she asserts. “One of the impacts of colonization on indigenous Africans, for example, is that children are often pushed out of schools because they can’t use the languages they’re learning — like Afrikaans — with their families at home.”
In fall 2025, she took a MISTI trip to Edinburgh, Scotland, where she studied sociolinguistics. She learned the value of considering alternative approaches to the brand of linguistics offered at MIT. “MIT’s approach to linguistics centers words and approaches its study like a math problem, while sociolinguistics includes important cultural context,” she says. Connecting the two made for a more complete, holistic approach to the work.
Honeycutt values a balanced approach to her studies, creating time for extracurricular activities that allow her to both investigate her research goals and create community. “I completed a policy internship in Washington, D.C. in 2024,” she recalls.
She’s a member of Theta Delta Chi, a fraternity comprising a diverse group of undergraduates from a variety of academic backgrounds. She plays women’s club soccer and is an officer with the MIT Undergraduate Association. As a co-chair of the Community Service committee, she’s leading efforts to create connections with students living off campus.
Honeycutt also volunteers with the Community Charter School of Cambridge, working to improve outcomes for underachieving students. As a volunteer, she’s able to pilot some of the education ideas being developed in her coursework. “I want to help underperforming students in the same way some institutions aid high-performing students,” she says.
The human element
Language shapes the ways its users view the world, according to Honeycutt. “I’m interested in how language can constrain thought,” she says. Language mastery is also a valuable tool in gauging emotional intelligence. “It’s important that people acquire and understand language in school,” she argues. “People should have access to a language that allows them to effectively communicate what they’re thinking.”
Having words for emotions can help people process them, Honeycutt believes. This is important in areas like translation and psychology, where nuance can be important. She also believes that reading and language acquisition are essential tools in developing effective self-awareness. Language is a medium for thought and provides guardrails to improve understanding.
“Access to a large vocabulary, including words for emotions, can increase your emotional intelligence,” she says.
With a solid academic foundation focused on cognition, language, and AI in place, Honeycutt plans to pursue studies in law and policy after graduation. That means law school and public policy programs, perhaps at an institution that offers a dual degree track.
“I want to extend opportunities to underserved students,” she says. “Problems in policy spaces are difficult, in part, because they defy easy categorization and involve multiple stakeholders.” Education, Honeycutt says, “is a fun problem to try to solve.” She wants to support efforts to enact lasting change by improving literacy, ensuring linguistic diversity, and centering science and research when crafting and implementing effective legislation that benefits learners, institutions, families, and communities.
There’s no single study in a field that will answer all the questions, Honeycutt argues. By combining the science of brain function with the social and mathematical aspects of linguistics, she can continue investigating language, its usage, and impacts on people and their lives. We can’t solve education challenges, improve AI and access to AI-enabled tools, and further the study of linguistics without institutional and community support.
“Support research,” Honeycutt says. “Don’t give up on trying to solve these problems.”
Unlocking mysteries of the universe through mathMathematician Amanda Burcroff is developing frameworks for understanding algebraic and geometric spaces in science as part of the School of Science Dean’s Postdoctoral Fellowship.GPS navigation, cryptography, quantum computing — while some of humankind’s greatest advancements have been invented by pioneers from various cultures, they were founded upon one common grammar: mathematics.
“Mathematics is the language with which God wrote the universe,” said the famous Italian astronomer, physicist, and philosopher Galileo Galilei, who, among his various scientific contributions, helped provide evidence for the idea that the sun is at the center of the solar system.
Although mostly conveyed through combinations of numbers, letters, and signs that may seem enigmatic to many, math equations hold within them countless stories — playbooks that generations of wonderers and inventors have crafted, refined, and shared in an attempt to make sense of a world full of unknown variables.
“I have faith in mathematics that, when there seems to be something special happening, when there’s some coincidence, that it’s not just a coincidence,” says mathematician Amanda Burcroff, “but that there’s actually some really deep, interesting, and involved reason for why that should be true.”
Burcroff’s research is focused on algebraic combinatorics, an area that provides discrete frameworks for understanding algebraic and geometric spaces that ubiquitously arise across science. This year, she joins MIT’s Department of Mathematics as a postdoc as part of the School of Science Dean’s Fellowship. Working with Professor Alexander Postnikov, Burcroff is building upon her techniques with the goal of applying them to other areas such as theoretical physics — a field that seeks to uncover the fundamental laws governing everything from subatomic particles to the cosmos itself.
“I have trust that if you keep following the path, eventually you’ll find the treasure — that is, whatever theorem or proof — that you’re looking for,” she says.
Exploring possibilities and redefining rules
Like many children, Burcroff once saw math as a subject that entailed lots of memorizing. Although she felt that it came naturally to her, she didn’t always find math very interesting.
In high school, as she came to learn about areas like calculus and geometry, Burcroff started to see the discipline in a different light — a creative approach to exploring what’s possible.
“[In] most other fields, the rules are imposed on you by the world,” she says, “but in math, you get full freedom to lay down those rules and then figure out what the implications of those rules are by using logical consequence.”
In 2015, Burcroff began her bachelor’s degree at the University of Michigan with a major in math and a minor in computer science. There, she entered the world of combinatorics — a branch of math dealing with counting, arranging, and combining objects that forms a crucial basis for understanding the complexity of problems, as well as the limits of computer algorithms.
“When I was starting out, I was just happy to have any mystery that anyone gave me,” she says.
Math was, to Burcroff, like a fun game with levels to complete. But during a study abroad program in Budapest, Hungary — the hometown of Paul Erdős, who is considered to be one of the most prolific mathematicians of the 20th century — it became more exciting to play when she was handed puzzles no one has yet solved.
“It turns out that if you put down the right set of rules, there’s an infinite number of beautiful things that you can do with it,” she says.
A journey of endless mysteries to unlock
In 2019, Burcroff embarked on a journey to pursue further research in England, later completing a master’s degree in pure mathematics at the University of Cambridge, then a research master’s degree at Durham University. In 2021, she returned to the United States and began her PhD at Harvard University, with the guidance of Professor Lauren Williams.
Among several riddles she has unraveled over the years, Burcroff helped unify different mathematical approaches to understand why systems work so reliably. Think of it as finding out that two seemingly different set of instructions actually lead the same way. By demonstrating their connections, her work has revealed an underlying, overarching mathematical architecture — a finding that later helped Burcroff and her collaborators tackle one of the many enduring riddles in her field.
Generalized cluster algebras form the basis for describing geometries that appear throughout physics. For more than a decade, mathematicians suspected these building blocks were created only by adding up ingredients and never subtracting, although no one was able to prove it. In 2024, Burcroff and her collaborators published a paper demonstrating that these spaces have nice positivity properties by developing a new way to count and organize patterns — helping untangle a long-standing conjecture, whose potential implications span from predicting particle collision outcomes to describing the spaces appearing in string theory.
These findings have earned Burcroff numerous prestigious awards including a National Science Foundation Graduate Research Fellowship, a British Marshall Scholarship, and a Jack Kent Cooke Graduate Fellowship.
Despite the tremendous number of problems she has answered, new ones keep arising.
“Every time you unlock one of them, it gives you a bunch of paths to new connected mysteries,” Burcroff says.
At MIT, she is working with Postnikov, whose research on combinatorics and positivity-type problems has presented a radically different way to calculate fundamental quantities in quantum field theory.
“Burcroff is conducting research across disciplinary boundaries,” says Postnikov.
He adds: “I am sure that she will have a lot of fruitful interactions with researchers in other MIT departments.”
Burcroff’s goal is to apply combinatorial techniques to broader physical contexts and direct applications, especially those with implications to topics like mirror symmetry, a principle in string theory suggesting that very different-looking geometric spaces can be mathematically equivalent.
While “doing math is 99 percent trying something and failing,” Burcroff says it is this same challenge that keeps her motivated. To her, it is not about reaching a destination, but rather about the continuous “process of discovery,” one she hopes to share beyond the typical classroom.
To make math more accessible, especially among underrepresented groups, Burcroff has worked with mentorship programs including Harvard’s Real Representations and Math Includes, Cambridge Girls’ Angle, and MIT PRIMES. During her time as a postdoc, she hopes to continue this outreach and explore ways to get involved with other support groups at MIT’s Department of Mathematics.
How neurons sense bacteria in the gutNeural interaction with bacteria has important effects on animal brains. A new study investigates how neurons sense bacteria by revealing, in nematodes, the bacterial signals that a key neuron detects.Recent studies suggest animals and people alike have close and complex relationships with the bacteria around and within them. The human gut microbiome, for instance, has been associated with both depression and Parkinson’s disease. To go beyond association toward understanding of the actual mechanisms that enable the bacterial microbiome to influence brain function, a new study by neuroscientists in The Picower Institute for Learning and Memory at MIT examines the mechanisms at work in a model “bacterial specialist,” the nematode Caenorhabditis elegans.
In the new open-access study in Current Biology, the team, led by Picower Fellow Cassi Estrem in the Picower Institute for Learning and Memory lab of Associate Professor Steven Flavell, identifies the specific chemicals that a key neuron in C. elegans senses, both in the bacteria that it eats and in the bacteria that it needs to avoid ingesting.
“In our bodies, our own cells are outnumbered by the bacterial cells living in and on us. There’s an increasing recognition that this has a profound impact on human health,” says Flavell, an investigator of the Howard Hughes Medical Institute and faculty member of MIT’s Department of Brain and Cognitive Sciences. “It’s been clear that there are links for some time. Our study aimed to identify the hard mechanisms of how a host nervous system is affected by bacteria in the alimentary canal.”
Achieving a fundamental mechanistic understanding of how neurons interact with bacteria could help improve attempts to intervene in or manipulate those interactions with therapeutic drugs or supplements, Flavell says.
Mmm … sugar
Flavell calls C. elegans a “bacterial specialist” because the tiny, transparent worm has evolved to eat bacteria as its diet, while also needing to avoid pathogenic bacteria that can prove to be its undoing. This has led it to develop a nervous system especially well-attuned to sorting out what is food and what is foe. In 2019, the lab discovered that the neuron NSM, which projects into the worm’s alimentary canal, employs two “acid sensing ion channels” (ASICs) to detect when certain bacteria have been ingested. Notably, those ion channels are analogous to ones found in neurons in humans. When NSM detects yummy bacteria, it releases serotonin that causes the worm to increase its feeding rate and slow its slithering so that it can stay to dine on the surrounding meal.
To really understand how this works, Flavell and Estrem realized they needed to know exactly what the ion channels are detecting in the bacteria. To get started, they exposed worms to 20 different kinds of bacteria the worms are known to encounter and found that they all activated NSM activity to varying extents. Then they broke the bacteria down into more and more specific chemical components to see which one or ones triggered NSM. The experiments ruled out many components, including DNA, lipids, proteins, and simple sugars, and instead found that it’s specifically the polysaccharide sugars that coat many bacteria that drive NSM activation. In particular, in gram-positive bacteria, a chemical called peptidoglycan activated NSM. In gram-negative bacteria, a different polysaccharide was apparently in play.
Estrem and Flavell’s team also ran experiments showing that polysaccharides from bacteria in general, and peptidoglycan in particular, not only trigger NSM electrical activity, but actually promote the feeding and slowing behaviors. They also showed that genetically knocking out the ASICs abolished these responses. In all, they demonstrated that polysaccharide and peptidoglycan detection are sufficient to trigger the worm’s behaviors, and requires the ASICs.
Better not eat this
Having shown what exactly triggers the worms to recognize their bacterial food, the researchers wondered whether they could also pinpoint a danger sign the worm finds in harmful bacteria. For these experiments, they carefully used Serratia marcescens, a bacterium that’s also infectious for humans. Some strains of the bacteria have a red color, while others do not. The red ones, which have a pigment called prodigiosin, tend to be much more lethal for worms. In their testing, the researchers found that when NSM detected the non-pigmented bacteria, the neuron still activated and the worms still ingested the bacteria, but when prodigiosin was present, NSM did not activate and the worm did not pump it in or slow down to eat.
Adding prodigiosin to normally yummy bacteria also suppressed NSM’s usual response. In other words, the worms have evolved their digestive behavior (and the detectors within NSM) to avoid ingesting a chemical specifically associated with danger.
Flavell says it’s likely that some of the fundamental mechanisms highlighted in the new paper will inform studies of similar mechanisms in other animals.
“We developed a way of identifying these pathways by studying this organism that specializes in bacterial detection and displays robust responses,” Flavell explains. “But there’s no reason these pathways should be limited to C. elegans. The molecular players we identified are found in many species, including mammals.”
In addition to Estrem and Flavell, the paper’s other authors are Malvika Dua, Colby Fees, Greg Hoeprich, Matthew Au, Bruce Goode, and Lingyi Deng.
The National Institutes of Health, the McKnight Foundation, the Alfred P. Sloan Foundation, the Howard Hughes Medical Institute, and The Freedom Together Foundation provided support for the study.
A materials scientist’s playgroundNew system at MIT.nano will support quantum technology research.Scientists and engineers around the world are working to improve quantum bits, or qubits, the minuscule building blocks of the quantum computer. Qubits are incredibly sensitive, making it easy for errors to be introduced, lowering device yield. But a new cluster tool at MIT.nano introduces capabilities that will allow researchers to continue advancements in qubit performance.
Passersby outside MIT.nano may have recently noticed a complex looking piece of equipment being installed on the first-floor cleanroom. What looks like a sci-fi movie prop is actually a state-of-the-art, custom-built molecular beam epitaxy (MBE): a physical vapor deposition system that operates under ultra-high vacuum to produce high-quality thin films. With the ability to grow different crystalline materials on a wafer, the tool will support quantum researchers and materials scientists by allowing them to study how film growth affects the properties of the materials used in making qubits.
“To realize the full promise of quantum computing, we need to build qubits that are robust, reproducible, and extensible,” says William D. Oliver, the Henry Ellis Warren (1894) Professor of Electrical Engineering and Computer Science and professor of physics at MIT. “To date, most of the improvements to superconducting qubit performance are traceable to circuit design — essentially, designing qubit circuits that are less sensitive to their environmental noise. However, those improvements have largely run their course. Going forward, we need to address the fundamental materials science and fabrication engineering required to reduce the sources of environmental noise. This multi-chamber, cassette-loaded, 200-millimeter wafer MBE system is exactly the right tool at the right time. And there’s no place better to do this research than at MIT.nano.”
That is because MIT.nano is preconditioned to receive this type of system with physical space, climate controls, policies and procedures for researchers, and expert staff to manage the lab. Through an equipment support plan, Oliver’s Engineering Quantum Systems (EQuS) group is able to install and run the tool inside MIT.nano, a high-performance, safe, and reliable environment.
A controlled environment is essential for the MBE. “Think of this system like an inverted International Space Station (ISS),” explains Patrick Strohbeen, research scientist in the EQuS group. “The ISS is a small chamber of atmosphere surrounded by the vacuum of space. This MBE system is a chamber of space-level vacuum surrounded by atmosphere.” That vacuum of space is kept at a steady negative 90 degrees Celsius, which enables precise growth of thin films on an atomic scale. It is the largest single deposition chamber (1-meter diameter) the manufacturer, DCA, has sold in the United States.
The journey of a wafer
The system, which in total takes up 600 square feet, is made up of six chambers. First is the load lock, where the wafer is placed into the system and brought down from atmospheric pressure to near the vacuum level of space. Then, the wafer enters the distribution center. This space acts like a central hub, transferring the wafers to other chambers. Next is the deposition, or “growth,” chamber. This is where the system’s primary function takes place — depositing materials, specifically atoms of superconducting metal, onto a substrate, typically silicon. From there, it moves to the oxidation chamber, which facilitates the growth of key ceramic materials for qubits. A fifth storage chamber can hold an additional 10 wafers within the vacuum.
A unique aspect of this system is its sixth chamber, designed for X-ray photoelectron spectroscopy (XPS). Using this chamber, researchers can shoot a photon in the form of X-rays at the surface and, when it hits the surface, it will excite the electron inside the material so that the electron jumps out and is picked up by a sensor that then tells the researcher about the environment the electron came from. As individual layers of atoms are put down in the growth chamber, scientists can move the wafer to the XPS chamber to measure changes in the material structure of the film and back again, all while keeping it inside the vacuum space.
Why is this important? “The quantum community has excellent device physicists and device engineers,” said Strohbeen. “The last piece of the puzzle is: We need to understand the materials platform that we’re using for these devices.” The buried interfaces, so far, have been understudied due to the difficulty in probing them, he explained.
For those of us who are not MBE experts, think of the snow that fell in Massachusetts this winter. How can you tell how much ice is on the pavement without removing all of the snow on top of it? And without changing the natural setting where the snow, ice, and pavement meet? With this system, specifically the XPS chamber, scientists can study the interfaces of buried materials without disturbing the physical or chemical environments. “It is a materials scientist’s playground,” jokes Strohbeen — a controlled space where researchers can learn about and explore materials’ interactions within layers of atoms.
Why MIT.nano?
When Oliver, who is also the director of the MIT Center for Quantum Engineering, secured the MBE Quantum, the next question was where to put it. Enter MIT.nano. Housing 45,000 square feet of cleanroom, this facility exists at MIT to support complex, sensitive equipment with both the infrastructure and the staff needed to maintain it.
“MIT.nano’s ultra-stable building utilities and lab environment are exactly what is needed to support a system that demands extreme repeatability and purity,” says Nick Menounos, MIT.nano associate director of infrastructure. “The success of this installation grew from the early collaboration. Professor Oliver engaged the MIT.nano team in the procurement process almost two years in advance. That foresight, combined with the infrastructure momentum we gained from the recent CHIPS Act project, meant that we could prepare the cleanroom perfectly. We compressed the installation process that normally takes several months and had this extraordinary machine running in under three weeks.”
“From the very beginning, the MIT.nano staff were helpful, knowledgeable, and willing to go above and beyond to make this happen,” says Oliver. “While the MIT.nano facility is certainly an infrastructural crown jewel at MIT, it’s the MIT.nano staff who make it the national treasure it is today.”
Positioning the MBE Quantum in the cleanroom helps the team focus on scalability and device yield. Humidity and particle count, two things carefully measured and maintained at MIT.nano, can affect the output of the device. Minimizing as many variables as possible is key to improving qubit performance. The cleanroom also allows for new device research because an array of fabrication and metrology tools are available without having to leave the clean environment.
“We’re really excited to see what we can do with it,” says Strohbeen. “We bought it as a materials science tool, and it will also be a device development tool due to the flexibility of having it in the cleanroom.”
The MBE system was purchased through a combination of grants from the Army Research Office (ARO) and from the Laboratory for Physical Sciences (LPS). The ARO grant, a Defense University Research Instrumentation Program grant, is the premier grant from ARO for funding large capital equipment purchases that should prove disruptive in technologically relevant areas. It arrives at an important time on campus, as one of MIT’s strategic initiatives — the MIT Quantum Initiative — aims to apply quantum breakthroughs to the most consequential challenges in science, technology, industry, and national security.
Transforming deep-space signals into cathedral soundAn immersive sound installation at Oulu Cathedral, Finland, co-created by MIT Associate Professor Kiyoshi Masui, transforms more than 4,000 cosmic signals into spatial audio.A new immersive sound installation at Oulu Cathedral, Finland, brings the research of MIT astrophysicist and associate professor of physics Kiyoshi Masui into a striking sensory form, transforming more than 4,000 cosmic signals into spatial audio.
With its grand opening on April 4, “The Logos” project invites visitors to experience deep-space phenomena not as distant abstractions, but as something immediate and resonant. The work is led by artist and creative technologist Andrew Melchior in collaboration with Masui, philosopher Timothy Morton, and cathedral dean Satu Saarinen. Together, they treat the cathedral, built in 1832, not just as a setting but as part of the instrument itself. Its stone surfaces and reverberant acoustics give physical presence to signals that have traveled from distant galaxies.
At the heart of the installation are data gathered by the Canadian Hydrogen Intensity Mapping Experiment (CHIME) radio telescope, which detects fast radio bursts (FRBs). FRBs are immensely energetic flashes lasting only milliseconds and originating in distant galaxies across the observable universe. The Logos represents one of the most extensive artistic sonifications of FRB data to date. Each day at noon, the cathedral is filled with a one-hour procedural composition derived from these bursts. Some bursts are singular events, never repeating, while others pulse again and again from unknown sources. These patterns remain one of astrophysics’ most compelling mysteries.
“The fast flashes will echo as snare-like beats bouncing through the cathedral,” says Masui. “The sweeping dispersion of the signal — where different radio frequencies arrive at slightly different times — creates harmonies between high and low tones. It should feel rich and layered, while also revealing something real about how these signals travel across billions of years of cosmic space before reaching Earth.”
By converting FRB data into a shared listening experience, the collaboration suggests a different way of understanding the universe: not only through analysis, but through attention.
Running through April 2027 to mark the cathedral’s 250th anniversary, The Logos will feature as part of Oulu2026 European Capital of Culture and the Lumo Art and Tech Festival.
Six from MIT awarded 2026 Paul and Daisy Soros Fellowships for New AmericansFellowship honors contributions of immigrants to American society by awarding $90,000 in funding for graduate studies.Six MIT affiliates — Denisse Córdova Carrizales SM ’26; Ria Das ’21, MNG ’22; Ronak Desai; Stacy Godfreey-Igwe ’22; Arya Rao; and Ananthan Sadagopan ’24 — have been named 2026 P.D. Soros Fellows. In addition, P.D. Soros Fellow Avinash Vadali will begin a PhD in condensed-matter physics at MIT this fall.
The fellowship provides immigrants and the children of immigrants up to $90,000 in tuition and stipend support for up to two years of graduate studies. Interested students should contact Kim Benard, associate dean of distinguished fellowships in Career Advising and Professional Development.
Denisse Córdova Carrizales
Córdova Carrizales SM '26 is a PhD student in nuclear science and engineering in the lab of Professor Mingda Li, where she completed her master's work earlier this year. She is working on synthesizing and characterizing quantum materials with the goal of bridging fundamental science and industry to make our technology more energy-efficient and sustainable.
Córdova Carrizales, who is of Mexican descent, grew up in Houston, Texas, before attending Harvard University, where she graduated in 2023 with a BA in physics. At Harvard, she dove into experimental condensed-matter research. She also conducted research with the Princeton Plasma Physics Laboratory, Commonwealth Fusion Systems, and VEIR, spanning computational plasma physics and high-temperature superconducting magnet and cable engineering.
Her work includes coauthored papers in Nature Physics, Nature Materials, and Advanced Materials, as well as lead-author publications in Nano Letters and Physical Review Materials. In 2023, she received the LeRoy Apker Award from the American Physical Society.
Beyond research, Córdova Carrizales has advocated in Congress for nuclear disarmament and risk reduction and has written a piece on the nuclear stockpile stewardship program. At Harvard, she founded an organization to support first-generation college students studying physics. In a completely different arena, she performed as the lead in an off-Broadway show in New York.
Ria Das
Das ’21, MNG ’22 is a PhD student in the MIT Department of Electrical Engineering and Computer Science. She graduated from MIT in 2021 with a BS dual degree in mathematics along with electrical engineering and computer science, and received her master of engineering degree in 2022.
The daughter of Indian immigrant parents, Das grew up in Nashua, New Hampshire, where she struggled with issues of belonging and identity. These questions came to the forefront during her PhD studies at Stanford University. Das decided to step off the academic treadmill by taking a leave from her PhD to think more deeply about these topics.
During her leave, she traveled around the country before moving to New York to work at Basis Research Institute, an AI research nonprofit. As a research associate, Das developed an urban data team that worked with federal and municipal government agencies on issues of economic and housing equity, blending her interests in science and social problems. She then returned to MIT to complete her doctoral studies.
Today, Das works with Professor Joshua Tenenbaum in the Department of Brain and Cognitive Sciences to study how people undergo conceptual change to build more robust, accessible systems for automated (social) science and improved educational design. Looking ahead, she hopes to become a professor, collaborating closely with policy practitioners.
Ronak Desai
Desai is currently a student in the Harvard/MIT MD-PhD program, where his PhD focuses on chemistry. The son of immigrants from Gujarat, India, Desai was born in Tyler, Texas, and grew up in nearby Lindale. He earned his undergraduate degree at the University of Texas at Austin.
Desai spent a semester interning at the U.S. House of Representatives as a Bill Archer Fellow. He also completed biomedical research focused on studying and engineering novel polyketide synthases, aspiring to produce next-generation antibiotics by harnessing such newly engineered synthases.
Desai graduated with degrees in chemistry and biochemistry as a first-generation college student, Health Science Scholar, and Dean’s Honored Graduate, receiving nine scholarships throughout college. His research has resulted in publications in journals such as Cell and Nature Communications.
Desai hopes to combine his passions for medicine, science, and public policy in his career to advance the treatment of infectious diseases. He is conducting his doctoral research under Professor James J. Collins in the MIT Department of Biological Engineering and the Harvard-MIT Program in Health Sciences and Technology. Desai’s research centers on using artificial intelligence to discover and design novel antibiotics, an opportunity to advance treatments for patients worldwide.
Stacy Godfreey-Igwe
Godfreey-Igwe ’22 attended MIT as a QuestBridge and Gates Scholar, graduating in 2022 with a BS in mechanical engineering and a concentration in sustainable design. A Burchard Scholar, she also became the first student at MIT to complete a major in African and African diaspora studies. After graduating, she pursued a science policy fellowship in Washington and interned at the U.S. Department of Energy’s Building Technologies Office, where she worked to broaden adoption of heat pump technologies across diverse stakeholders.
Growing up in Richardson, Texas, as the daughter of Nigerian immigrants, Godfreey-Igwe developed an early awareness of structural inequality, particularly in how families like hers managed the burden of the severe Texas heat and high electricity costs. These experiences formed the basis of her lifelong journey seeking to address systemic inequities embedded in everyday systems.
Godfreey-Igwe is currently a doctoral student in the joint engineering and public policy - civil and environmental engineering program at Carnegie Mellon University (CMU), where she was selected for the inaugural CMU Rales Fellowship cohort. At CMU, she studies the impact of extreme heat on household energy use, particularly in vulnerable communities.
Beyond her research, Godfreey-Igwe organizes outreach and programming for local underrepresented students in STEM and participates in institutional efforts to expand access and belonging among graduate students. She aims to be a scholar and advocate whose work, drawing on her personal experiences, informs equitable energy solutions in a warming world.
Arya Rao
Rao is a student in the Harvard/MIT MD-PhD program. She completed her undergraduate degrees in biochemistry and computer science at Columbia University. Working with professors Pardis Sabeti (Harvard University) and Sangeeta Bhatia (MIT), Rao uses evolution as a lens for therapeutic design, developing artificial intelligence methods that read the genetic record and guide new intervention strategies.
Leveraging her dual training in medicine and computer science, Rao also leads the MESH AI Research Group at Mass General Brigham, where she develops simulation-based tools that test clinical AI systems in realistic educational settings before they reach patients.
Rao has been recognized for her work with a Forbes 30 Under 30 honor, the Massachusetts Medical Society Information Technology Award, the Harvard Presidential Public Service Fellowship, a Harvard Medical School Dean’s Innovation Award, and a Ladders to Cures Accelerator Award. She has published more than 30 manuscripts in publications including JAMA, Nature, and NEJM AI.
Growing up in rural northern Michigan, Rao was inspired by her parents, Konkani immigrants from India, who served as two of the area’s only physicians. She has always imagined a career that could leverage scientific innovation to improve patient care, especially for communities without access like her own. Going forward, she envisions a career as a surgeon-scientist that keeps her close to patients while taking on leadership that shapes how new technologies are evaluated, implemented, and made usable in the places that need them most.
Ananthan Sadagopan
Sadagopan ’24 grew up in Westborough, Massachusetts, as the child of immigrants from Chennai, India. He participated in chemistry competitions, winning the You Be the Chemist Challenge in middle school and earning a gold medal at the International Chemistry Olympiad for the United States in high school. He attended MIT for college, graduating in three years in 2024 with a bachelor’s degree in chemistry and biology.
At MIT, Sadagopan worked with Srinivas Viswanathan on computational biology projects and with William Gibson, Matthew Meyerson, and Stuart Schreiber on chemical biology projects. He led projects characterizing somatic perturbations of X chromosome inactivation in cancer, developing a machine-learning tool for cancer dependency prediction, using small molecules to relocalize proteins in cells, and creating a generalizable strategy to drug the most mutated gene in cancer, TP53. Sadagopan’s work has been patented and published in journals such as Cell and Nature Chemical Biology.
Sadogopan was president of the chemistry undergraduate association and led the events committee for MIT Science Olympiad. He is currently pursuing a PhD in biological and biomedical science at Harvard University as a Hertz Fellow and Herchel Smith Fellow. He is interested in de-risking new therapeutic strategies and hopes that his work will inspire pharma companies to bring first-in-class therapies to patients.
Three from MIT named 2026 Goldwater ScholarsRising seniors Deeksha Kumaresh, Anna Liu, and Charlotte Myers are honored for their academic achievements.Three MIT rising seniors have been selected to receive a 2026 Barry Goldwater Scholarship, including Deeksha Kumaresh in the School of Engineering and Anna Liu and Charlotte Myersin the School of Science. An estimated 5,000 college sophomores and juniors from across the United States were nominated for the scholarships, of whom only 454 were selected.
The Goldwater Scholarships have been conferred since 1989 by the Barry Goldwater Scholarship and Excellence in Education Foundation. These scholarships have supported undergraduates who go on to become leading scientists, engineers, and mathematicians in their respective fields.
Deeksha Kumaresh, a third-year biological engineering major, is an undergraduate researcher at the Hammond Lab. The Hammond Research Group at the MIT Koch Institute for Integrative Cancer Research focuses on the self-assembly of polymeric nanomaterials, with a major emphasis on the use of electrostatics and other complementary interactions to generate multifunctional materials with highly controlled architecture.
“Hands down, the mentors I’ve encountered have been the most significant part of my MIT journey,” Kumaresh says. “I’m also extremely grateful to the Hammond Lab, which has provided a supportive environment where I can make mistakes, learn, and grow as a researcher. I treasure the spontaneous conversations with lab members (about science or life) and their willingness to treat me seriously as an independent researcher, even as an undergraduate.”
Kumaresh is mentored by Paula Hammond, dean of the School of Engineering, Institute Professor, and professor of chemical engineering. Kumaresh's career goals are to pursue an MD/PhD. In the long term, she seeks to lead a bioengineering research lab to predict the efficacy and side effects of cancer therapies by developing systems-level computational and biological preclinical models.
“Receiving this scholarship has been incredibly meaningful, because it offered me the chance to reflect critically on my post-graduate goals and receive recognition for my journey for them,” Kumaresh says. “Earning this scholarship has welcomed me into a tight-knit community where I’ve already found so much guidance. Everyone is genuinely curious about everyone else’s interests and are eager to lend a hand however they can.”
Anna Liu, a third-year chemistry major, is an undergraduate researcher in the Radosevich Group. The overarching objective of the group’s research is to develop new catalysts, strategies, and reagents for synthetic chemistry. By designing and synthesizing new molecular compounds with unknown structure and function, the group hopes to learn more about the general principles enabling new chemical transformations.
Liu is mentored by professor of chemistry Alexander Radosevich. She plans to pursue a PhD in organic or inorganic chemistry and eventually lead research developing sustainable synthetic transformations informed by fundamental mechanistic and reactivity studies, and teach at the university level.
“Going through the Goldwater application process gave me a deeper understanding of my research project and helped me reflect on my intrinsic motivations to pursue research. I’m excited to use what I’ve learned to keep growing as a researcher,” Liu says. “I am so grateful for the countless mentors, teachers, labmates, classmates, friends, and family in my life who have believed in me, fostered my passion for chemistry, and taught me so much. Receiving this scholarship is truly a testament to their outstanding support!"
Charlotte Myers, a third-year physics and astronomy major, conducts research at the Kavli Institute for Astrophysics and Space Research, where she applies machine learning to model galactic structure, and at the Center for Theoretical Physics, where she studies theoretical models of dark matter. Her research interests center on the physics of dark matter, which she approaches from multiple perspectives — from its distribution on galactic scales to particle-level models.
Myers is mentored by Lina Necib, an assistant professor in the Department of Physics. She plans to pursue a PhD in theoretical physics and conduct research in cosmology and astroparticle physics, with a focus on the fundamental physics of dark matter, and teach at the university level.
“I am very grateful to my research advisors, Professor Necib, Dr. Starkman, and Professor Slatyer, for their guidance and support in helping me develop as a researcher,” Myers says. “I find it deeply rewarding to engage with open questions in physics, and I am excited to continue pursuing this work in graduate school and beyond. Receiving this scholarship has given me both the resources and the confidence to continue on that path, even when progress is not always linear.”
The scholarship program honoring Senator Barry Goldwater was designed to identify, encourage, and financially support outstanding undergraduates interested in pursuing research careers in the sciences, engineering, and mathematics. The Goldwater Scholarship is the preeminent undergraduate award of its type in these fields.
MIT takes top team honors in 86th Putnam Math CompetitionThe undergraduate team topped the scoreboard for the sixth year in a row and also took the Elizabeth Lowell Putnam Prize again.In an outstanding performance at the 86th William Lowell Putnam Mathematical Competition, MIT’s team once again took the top spot for the sixth consecutive year. MIT secured four of the five Putnam Fellows, who are the five highest-ranking students, and the Elizabeth Lowell Putnam Prize, which is given to a woman whose “performance in the competition is particularly meritorious.”
The members of the winning team, consisting of junior Cheng Jiang, senior Luke Robitaille, and first-year Chunji Wang, were all awarded as Putnam Fellows alongside senior Zixiang Zhou, each receiving a $2,500 award for their performance. Notably, Robitaille is a four-time Putnam Fellow, having received the award for each year of his studies. For a second consecutive year, sophomore Jessica Wan was awarded the Elizabeth Lowell Putnam Prize and received $1,000.
Wan was also among the top 25 scorers, amongst 16 others from MIT: Warren Bei, Reagan Choi, Pico Gilman, Henry Jiang, Zhicheng Jiang, Papon Lapate, Gyudong Lee, Derek Liu, Maximus Lu, Krishna Pothapragada, Pitchayut Saengrungkongka, Qiao Sun, Allen Wang, Kevin Wang, and Yichen Xiao.
A legacy of success
“I was delighted to see how well the MIT students did on the Putnam exam this year, which reflects their hard work, talent, and enthusiasm,” says Professor Henry Cohn, who led class 18.A34 (Mathematical Problem Solving) this year, also informally known as the Putnam seminar.
MIT’s continued success in the Putnam competition stems from a variety of sources. Some of this is built on things like the seminar, where students get together to sharpen their skills by diving deep into tough problems and discussing solutions.
Cohn, a former participant in the Putnam, comments on the joy of teaching the seminar and seeing students’ progress. “When you spend a semester watching students present solutions to difficult problems, you start to understand how they think,” says Cohn. “It’s exciting to see them apply their abilities to new, difficult problems."
Professor Bjorn Poonen, who also led the seminar in previous years (and is a four-time Putnam Fellow), describes it as an opportunity to hone a spectrum of skills in competition preparation. “Knowing how to explain things well is really important for doing well on the Putnam and for everything else, and for this it really helps to have experience communicating with others, which is what the problem-solving seminar is all about.”
A shared passion for problem-solving
The students who take the Putnam thrive on all aspects of the competition, from the social to the exam itself.
“It’s not a school day, and we still get to do math,” Jiang describes his excitement for the competition. Indeed, getting to “do math” extends beyond formally sitting for the exam, to breaks and opportunities for discussion that are interspersed throughout the day. The students take each opportunity to come together as seriously as they do the competition, and it is this collective passion for problem-solving that builds a strong sense of community and brings students back year after year.
“The competition brings together hundreds of students from across campus representing many majors, years of graduation, and degrees of math contest experience, but what brings everyone together is a shared love of solving problems,” Cohn says. “You can see this in the clusters of students who stay to discuss the problems long after the exam has ended. Mathematics can sometimes feel like a solitary pursuit, but at this level, collaboration is key.”
Community complements the shared passion the math enthusiasts share for problems and puzzles. “You get a kind of satisfaction similar to when you get unstuck while doing a crossword puzzle and everything falls into place,” Poonen describes his own experience solving Putnam problems.
Consistency in certainty
The competition is also an opportunity to see familiar faces. Robitaille recalls his experiences in high school math olympiads, and highlights the friendly atmosphere at the Putnam. “Throughout college, I have stayed close with people I met at competitions,” Robitaille says. “There’s the whole background of times spent together, not just on contest day.”
An event for both community and challenge, the consistency and certainty of competition day is what brought Robitaille and Zhou back year after year. “Each time, you have a set amount of time to sit in the room and work on the problems,” Robitaille says. “If you were the type of person for whom that would be a fun thing, like me, it’s nice to have an opportunity to do it again occasionally.”
“It’s more fun than the real world, where everything is complicated,” Zhou adds with a smile.
The full list of 2025 winners can be found on the Putnam website.
MIT affiliates elected to the American Academy of Arts and Sciences for 2026The prestigious honor society honors four MIT faculty and 13 additional MIT alumni among more than 250 new members.Four MIT faculty members are among the roughly 250 leaders from academia, the arts, industry, public policy, and research elected to the American Academy of Arts and Sciences, the academy announced April 22. Thirteen additional MIT alumni were also honored.
One of the nation’s most prestigious honorary societies, the academy is also a leading center for independent policy research. Members contribute to academy publications, as well as studies of science and technology policy, energy and global security, social policy and American institutions, the humanities and culture, and education.
MIT faculty elected from MIT in 2026 are:
MIT alumni elected this year include Mark Aguiar PhD ’99 (Economics); Mark G. Allen SM ’86, PhD ’89 (Chemical Engineering); Magdalena Balazinska PhD ’06 (EECS); Keren Bergman SM ’91, PhD ’94 (EECS); Sara Cherry PhD ’00 (Biology); Cynthia J. Ebinger SM ’86, PhD ’88 (EAPS); Charles L. Epstein ’78 (Mathematics); Shanhui Fan PhD ’97 (Physics); Atif Mian ’96, PhD ’01 (Mathematics with Computer Science and Economics); Sarah E. O'Connor PhD ’01 (Chemistry); Darryll J. Pines SM ’88, PhD ’92 (Mechanical Engineering); Phillip (Terry) Ragon ’72 (Physics); and Mansour Shayegan ’79, EE ’81, SM ’81, PhD ’83 (Electrical Engineering).
“We celebrate the achievement of each new member and the collective breadth and depth of their excellence – this is a fitting commemoration of the nation’s 250th anniversary,” said Academy President Laurie Patton.
Since its founding in 1780, the academy has elected leading thinkers from each generation, including George Washington and Benjamin Franklin in the 18th century, Maria Mitchell and Daniel Webster in the 19th century, and Toni Morrison and Albert Einstein in the 20th century. The current membership includes more than 250 Nobel and Pulitzer Prize winners.
Two MIT alumnae named 2026 Gates Cambridge ScholarsMitali Chowdhury ’24 and Christina Kim ’24 will pursue graduate studies at Cambridge University in the UK.Mitali Chowdhury ’24 and Christina Kim ’24 have been selected as 2026 Gates Cambridge Scholars. The highly competitive fellowship offers fully funded opportunities for postgraduate study in any field at Cambridge University in the U.K. Kim is a second-time Gates Cambridge Scholar.
MIT students interested in the Gates Cambridge Scholar program should contact Kim Benard, associate dean of distinguished fellowships in Career Advising and Professional Development.
Mitali Chowdhury
Chowdhury graduated from MIT with a BS in biological engineering and minors in both urban planning and environment and sustainability. Chowdhury has had a longstanding interest in reducing inequities in global health. At MIT, she pursued research in point-of-care diagnostics to identify and treat disease with accessible biotechnologies. She also helped develop low-cost testing for bacterial contamination in water in South Asia.
Chowdhury currently works at a startup advancing sequencing-based diagnostics. At Cambridge University, she will study for MPhil and PhD degrees in the Centre for Doctoral Training in Sensor Technologies. Her research will focus on CRISPR-based diagnostics to address antimicrobial resistance and expand equitable access to care.
Christina Kim
After graduating from MIT with a bachelor’s degree in chemistry and biology, Kim worked as a researcher in women’s health at the Wellcome Sanger Institute in Cambridge, U.K.
As a 2025 Gates Cambridge Scholar, Kim pursued an MPhil in research at the institute, focusing on using bioinformatics and tissue engineering to design novel in vitro models. Her second Gates Cambridge scholarship will fund her PhD studies.
Professor Michael Laub and MIT alumni named 2025 AAAS FellowsThe American Association for the Advancement of Science recognized Laub and 21 alumni for their efforts to advance science and related fields.MIT Professor Michael T. Laub as well as 21 MIT alumni have been elected as fellows of the American Association for the Advancement of Science (AAAS).
The 2025 class of AAAS Fellows includes 449 scientists, engineers, and innovators, spanning all 24 of AAAS disciplinary sections, who are recognized for their scientific achievements.
Laub, the Salvador E. Luria Professor in the MIT Department of Biology and an HHMI Investigator, studies the biological mechanisms and evolution of how cells process information to regulate their own growth and proliferation, using bacteria as a model organism to develop a deeper, fundamental understanding of how bacteria function and evolve. Laub was honored as a AAAS Fellow for distinguished contributions to the field of bacterial information processing, particularly to the understanding of coevolution of host-pathogen response and immunity.
“This year’s AAAS Fellows have demonstrated research excellence, made notable contributions to advance science, and delivered important services to their communities,” said Sudip S. Parikh, AAAS chief executive officer and executive publisher of the Science family of journals. “These fellows and their accomplishments validate the importance of investing in science and technology for the benefit of all.”
The following alumni were also named fellows of the AAAS:
MIT Associate Professor Jacob Andreas of the Department of Electrical Engineering and Computer Science [EECS] and MIT Associate Professor Brett McGuire of the Department of Chemistry have been selected as the winners of the 2026 Harold E. Edgerton Faculty Achievement Award. Established in 1982 as a permanent tribute to Institute Professor Emeritus Harold E. Edgerton’s great and enduring support for younger faculty members, this award is given annually in recognition of exceptional distinction in teaching, research, and service.
“The Department of Chemistry is extremely delighted to see Brett recognized for science that has changed how we think about carbon in space,” says Class of 1942 Professor of Chemistry and Department Head Matthew D. Shoulders. “Brett’s lab combines laboratory spectroscopy, radio astronomy, and sophisticated signal-analysis methods to pull definitive molecular fingerprints out of extraordinarily faint data. His discovery of polycyclic aromatic hydrocarbons in the cold interstellar medium has opened a powerful new window on astrochemistry. Moreover, Brett is inventing the creative and unique tools that make discoveries like this possible.”
“Jacob Andreas represents the very best of MIT EECS” says Asu Ozdaglar, EECS department head. “He is an innovative researcher whose work combines computational and linguistically informed approaches to build foundations of language learning. He is an extraordinary educator who has brought these forefront ideas into our core classes in natural language processing and machine learning. His ability to bridge foundational theory with real-world impact, while also advancing the social and ethical dimensions of computing, makes him truly deserving of the Edgerton Faculty Achievement Award.”
Andreas joined the MIT faculty in July 2019, and is affiliated with the Computer Science and Artificial Intelligence Laboratory. His work is in natural language processing (NLP), and more broadly in AI. He aims to understand the computational foundations of language learning, and to build intelligent systems that can learn from human guidance. Among other honors, Andreas has received Samsung’s AI Researcher of the Year award, MIT’s Kolokotrones and Junior Bose teaching awards, a 2024 Sloan Research Fellow award, and paper awards at the International Conference on Machine Learning and the Association for Computational Linguistics.
Andreas received his BS from Columbia University, his MPhil from Cambridge University (where he studied as a Churchill scholar), and his PhD in natural language processing from the University of California at Berkeley. His work in natural language processing has taken on thorny problems in the capability gap between humans and computers. “The defining feature of human language use is our capacity for compositional generalization,” explains Antonio Torralba, Delta Electronics Professor and faculty head of Artificial Intelligence and Decision-Making in the Department of EECS. “Many of the core challenges in natural language processing is addressed by simply training larger and larger neural models, but this kind of compositional generalization remains a persistent difficulty, and without the ability to generalize compositionally, the deep learning toolkit will never be robust enough for the most challenging real-world NLP tasks. Jacob’s work on compositional modeling draws new connections between NLP and work in computer vision and physics aimed at modeling systems governed by symmetries and other algebraic structures and, using them, they have been able to build NLP models exhibiting a number of new, human-like language acquisition behaviors, including one-shot word learning, learning via mutual exclusivity constraints, and learning of grammatical rules in extremely low-resource settings.”
Within EECS, Andreas has developed multiple advanced courses in natural language processing, as well as new exercises designed to get students to grapple with important social and ethical considerations in machine learning deployment. “Jacob has taken a leading role in completely modernizing and extending our course offerings in natural language processing,” says award nominator Leslie Pack Kaelbling, Panasonic Professor in the Department of EECS. “He has led the development of a modern two-course sequence, which is a cornerstone of the new AI+D [artificial intelligence and decision-making] major, routinely enrolling several hundred students each semester. His command of the area is broad and deep, and his classes integrate classical structural understanding of language with the most modern learning-based approaches. He has put MIT EECS on the worldwide map as a place to study natural language at every level.”
Brett McGuire joined the MIT faculty in 2020 and was promoted to associate professor in 2025. His research operates at the intersection of physical chemistry, molecular spectroscopy, and observational astrophysics, where he seeks to uncover how the chemical building blocks of life evolve alongside and help shape the birth of stars and planets. A former Jansky Fellow and then Hubble Postdoctoral Fellow at the National Radio Astronomy Observatory, McGuire has a BS in chemistry from the University of Illinois and a PhD in physical chemistry from Caltech. His honors include a 2026 Sloan Fellowship, the Beckman Young Investigator Award, the Helen B. Warner Prize for Astronomy, and the MIT Award for Teaching with Digital Technology.
The faculty who nominated McGuire for this award praised his extraordinary public outreach, his immediate willingness to take on teaching class 5.111 (Principles of Chemical Science), a General Institute Requirement (GIR) course comprised of 150–500 students, and his service to both the MIT and astrochemical communities.
“Brett is at the very top of astrochemical scientists in his age group due to his discovery of fused carbon ring compounds in the cold region of the ISM [interstellar medium], an observation that provides a route for carbon incorporation in planets,” says Sylvia Ceyer, the John C. Sheehan Professor of Chemistry in her nomination statement. “His extensive involvement in service-oriented activities within the astrochemical/physical community is highly unusual for a junior scientist, and is testament to the value that the astronomical community places in his wisdom and judgement. His phenomenal organizational skills have made his contributions to graduate admission protocols and seminar administration at MIT the envy of the department. And most importantly, Brett is a superb teacher, who cares deeply about students’ understanding and success, not only in his course, but in their future endeavors.”
“As an assistant professor, Brett volunteered to teach 5.111, a large GIR course with 150–500 students, and has received some of the best teaching evaluations among all faculty who have led the subject,” says Mei Hong, the David A. Leighty Professor of Chemistry. “He has a natural talent in explaining abstract physical chemistry concepts in an engaging manner. His slides, which he prepared from scratch instead of modifying from previous years’ material from other professors, are clear, and … the combination of lucid explanation and humor has generated great enthusiasm and interest in chemistry among students.”
Subject evaluations from McGuire’s courses praised his humor, the clarity of his explanations, and his ability to transform a lecture into a “science show.” “I haven't felt this sort of desire for the depth of understanding in a subject beyond just a straight grade [in some time],” says one student. “Brett definitely stimulated that love of learning for me.”
“Brett is an outstanding faculty member who is dedicated to fostering student learning and success,” says Jennifer Weisman, assistant director of academic programs in chemistry. “He is thoughtful, caring, and goes above and beyond to help his colleagues, students, and staff.”
“I’m thrilled to be selected for the Edgerton Award this year,” says McGuire. “The award is nominally for teaching, research, and service; MIT and the chemistry department in particular have been an incredible place to learn and grow in all these areas. I’m incredibly grateful for the mentorship, enthusiasm, and support I have received from my colleagues, from my students both in the lab and in the classroom, and from the MIT community during my time here. I look forward to many more years of exciting discovery together with this one-of-a-kind community.”
With navigating nematodes, scientists map out how brains implement behaviorsMIT scientists create a detailed map of exactly what happens in the brains of C. elegans worms when they “follow their nose” to savor attractive odors or avoid unappealing ones.Animal behavior reflects a complex interplay between an animal’s brain and its sensory surroundings. Only rarely have scientists been able to discern how actions emerge from this interaction. A new open-access study in Nature Neuroscience by researchers in The Picower Institute for Learning and Memory at MIT offers one example by revealing how circuits of neurons within C. elegans nematode worms respond to odors and generate movement as they pursue of smells they like and evade ones they don’t.
“Across the animal kingdom, there are just so many remarkable behaviors,” says study senior author Steven Flavell, associate professor in the Picower Institute and MIT’s Department of Brain and Cognitive Sciences and an investigator of the Howard Hughes Medical Institute. “With modern neuroscience tools, we are finally gaining the ability to map their mechanistic underpinnings.”
By the end of the study, which former graduate student Talya Kramer PhD ’25 led as her doctoral thesis research, the team was able to show exactly which neurons in the worm’s brain did which of the jobs needed to sense where smells were coming from, plan turns toward or away from them, shift to reverse (like old-fashioned radio-controlled cars, C. elegans worms turn in reverse), execute the turns, and then go back to moving forward. Not only did the study reveal the sequence and each neuron’s role in it, but it also demonstrated that worms are more skillful and intentional in these actions than perhaps they’ve received credit for. And finally, the study demonstrated that it’s all coordinated by the neuromodulatory chemical tyramine.
“One thing that really excited us about this study is that we were able to see what a sensorimotor arc looks like at the scale of a whole nervous system: all the bits and pieces, from responses to the sensory cue until the behavioral response is implemented,” Flavell says.
Seeing the sequence
To do the research, Kramer put worms in dishes with spots of odors they’d either want to navigate toward or slither away from. With the lab’s custom microscopes and software, she and her co-authors could track how the worms navigated and all the electrical activity of more than 100 neurons in their brains during those behaviors (the worms only have 302 neurons total).
The surveillance enabled Kramer, Flavell, and their colleagues to observe that the worms weren’t just ambling randomly until they happened to get where they’d want to be. Instead, the worms would execute turns with advantageous timing and at well-chosen angles. The worms seemed to know what they were doing as they navigated along the gradients of the odors.
Inside their heads, patterns of electrical activity among a cohort of 10 neurons (indicated by flashing green light tied to the flux of calcium ions in the cells), revealed the sequence of neural activation that enabled the worms to execute these sensible sensory-guided motions: forward, then into reverse, then into the turn, and then back to forward. Particular neurons guided each of these steps, including detecting the odors, planning the turn, switching into reverse, and then executing the turns.
A couple of neurons stood out as key gears in the sequence. A neuron called SAA proved pivotal for integrating odor detection with planning movement, as its activity predicted the direction of the eventual turn. Several neurons were flexible enough to show different activity patterns depending on factors such as where the odors were and whether the worm was moving forward or in reverse.
And if the neurons are indeed turning and shifting gears, then the neuromodulator tyramine (the worm analog of norepinephrine) was the signal essential to switch their gears. After the worms started moving in reverse, tyramine from the neuron RIM enabled other neurons in the sequence to change their activity appropriately to execute the turns. In several experiments the scientists knocked out RIM tyramine and saw that the navigation behaviors and the sequence of neural activity largely fell apart.
“The neuromodulator tyramine plays a central role in organizing these sequential brain activity patterns,” Flavell says.
In addition to Flavell and Kramer, the paper’s other authors are Flossie Wan, Sara Pugliese, Adam Atanas, Sreeparna Pradhan, Alex Hiser, Lillie Godinez, Jinyue Luo, Eric Bueno, and Thomas Felt.
A MathWorks Science Fellowship, the National Institutes of Health, the National Science Foundation, The McKnight Foundation, The Alfred P. Sloan Foundation, the Freedom Together Foundation, and HHMI provided funding to support the work.
A regulatory loophole could delay ozone recovery by yearsScientists say an exception in the Montreal Protocol for the use of ozone-depleting feedstocks could set the ozone recovery back seven years.Often hailed as the most successful international environmental agreement of all time, the 1987 Montreal Protocol continues to successfully phase out the global production of chemicals that were creating a growing hole in the ozone layer, causing skin cancer and other adverse health effects.
MIT-led studies have since shown the subsequent reduction in ozone-depleting substances is helping stratospheric ozone to recover. (It could return to 1980 levels by as early as 2040, according to some estimates.) But the Montreal Protocol made an exception in its rules for the use of ozone-depleting substances as feedstocks in the production of other materials. That’s because it was thought that only a small amount — just 0.5 percent — of the ozone-depleting substances used for this purpose would leak into the atmosphere.
In recent years, however, scientists have observed more ozone-depleting substances in the atmosphere than expected, and have increased their estimates of leakage from feedstocks.
Now an international group of scientists, including researchers from MIT, has calculated the impact of different feedstock leakage rates on the ozone’s fragile recovery. They find the higher leakage rates, if not addressed by the Montreal Protocol, could delay ozone recovery by about seven years.
“We’ve realized in the last few years that these feedstock chemicals are a bug in the system,” says author Susan Solomon, the Lee and Geraldine Martin Professor of Environmental Studies and Chemistry, who was part of the original research team that linked the chemicals to the ozone hole. “Production of ozone-depleting substances has pretty much ceased around the world except for this one use, which is when you have a chemical you convert into something else.”
The paper, which was published in Nature Communications today, is the first to comprehensively quantify the impact of leaked feedstocks, which are currently used to make plastics and nonstick chemicals. They are also used to make substitute chemicals for the ones regulated under the Montreal Protocol. The researchers say it shows the importance of curbing use and preventing leakage of such feedstocks, especially as the production of their end products, like plastic, is projected to grow.
“We’ve gotten to the point where, if we want the protocol to be as successful in the future as it has been in the past, the parties really need to think about how to tighten up the emissions of these industrial processes,” says first author Stefan Reimann of the Swiss Federal Laboratories for Materials Science and Technology.
“To me, it’s only fair, because so many other things have already been completely discontinued. So why should this exemption exist if it’s going to be damaging?” says Solomon.
Joining Reimann on the paper are his colleagues Martin K. Vollmer and Lukas Emmenegger; Luke Western and Susan Solomon of the MIT Center for Sustainability Science and Strategy and the Department of Earth, Atmospheric and Planetary Sciences; David Sherry of Nolan-Sherry and Associates Ltd; Megan Lickley of Georgetown University; Lambert Kuijpers of the A/gent Consultancy b.v.; Stephen A. Montzka and John Daniel of the National Oceanic and Atmospheric Administration; Matthew Rigby of the University of Bristol; Guus J.M. Velders of Utrecht University; Qing Liang of the NASA Goddard Space Flight Center; and Sunyoung Park of Kyungpook National University.
Repairing the ozone
In 1985, scientists discovered a growing hole in the ozone layer over Antarctica that was allowing more of the sun’s harmful ultraviolet radiation to reach Earth’s surface. The following year, researchers including Solomon traveled to Antarctica and discovered the cause of the ozone deterioration: a class of chemicals called chlorofluorocarbons, or CFCs, which were then used in refrigeration, air conditioning, and aerosols.
The revelations led to the Montreal Protocol, an international treaty involving 197 countries and the European Union restricting the use of CFCs. The subsequent decision to exempt the use of ozone-depleting substances for use as feedstocks was based partially on industry estimates of how much of their feedstocks leaked.
“It was thought that the emissions of these substances as a feedstock were minor compared to things like refrigerants and foams,” Western says. “It was also believed that leakage from these sources was minor — around half a percent of what went in — because people would essentially be leaking their profits if their feedstocks were released into the atmosphere.”
Unfortunately, some of those assumptions are no longer true. Western and Reimann are part of the Advanced Global Atmospheric Gases Experiment (AGAGE), a global monitoring network co-founded by Ronald Prinn, MIT’s TEPCO Professor of Atmospheric Science. AGAGE monitors emissions of ozone-depleting substances around the world, and in recent years researchers have revised their estimates of feedstock leakage upwards, to about 3.6 percent. For some chemicals, the number was even higher.
In the new paper, the researchers estimated a 3.6 percent feedstock leakage as the baseline for most chemicals. They compared that with a scenario where 0.5 percent of feedstocks are leaked from 2025 onward and a scenario with zero feedstock-related emissions. The researchers also looked at production trends between 2014 and 2024 to project how much of each specific ozone-depleting chemical would be used as feedstock between 2025 and 2100.
The analysis shows that until 2050, total ozone-depleting chemical emissions decrease in all scenarios as rising feedstock emissions are offset by declining uses enforced by the Montreal Protocol. In the scenario with continued 3.6 percent leakage, however, emissions level off around 2045, and total emissions only decrease by 50 percent overall by 2100.
The researchers then evaluated the impact of feedstock-related emissions on stratospheric ozone depletion. In the scenario where feedstock leakage is 0.5 percent, the ozone returns to its 1980 status by 2066. In the scenario with zero feedstock leakage, the ozone reclaims its 1980 health in 2065. But in the baseline scenario, the recovery is delayed about seven years, to 2073.
“This paper sends an important message that these emissions are too high and we have to find a way to reduce them,” Reimann says. “Either that means no longer using these substances as feedstocks, swapping out chemicals, or reducing the leakage emissions when they are used.”
A global response
Solomon is confident industries will be able to adjust to the latest findings.
“There are a lot of innovators in the chemical industry,” Solomon says. “They make new chemicals and improve chemicals for a living. It’s true they can perhaps get too entrenched with certain chemicals, but it doesn’t happen that often. Actually, they’re usually quite willing to consider alternatives. There are thousands of other chemicals that could be used instead, so why not switch? That’s been the attitude.”
Solomon says the fact that AGAGE can detect the impact of feedstock emissions is a testament to the progress the world has made in reducing emissions from other sources up to this point. She believes raising awareness of the feedstock problem is the first step.
“This isn’t the first time that the AGAGE Network has made measurements that have allowed the world to see we need to do a little better here or there,” Western says. “Often, it’s just a mistake. Sometimes all it takes is making people more aware of these things to tighten up some processes.”
Members of the Montreal Protocol meet every year. In those meetings, they split into working groups around different topics. Feedstock emissions are already one of those topics, so participants will review the evidence together. Typically, they release a statement about mitigation strategies if needed.
“We wanted to raise the warning flag that something is wrong here,” Reimann says. “We could reduce the period of ozone depletion by years. It might not sound like a long time, but if you could count the skin cancer cases you’d avoid in that time, it would seem quite significant.”
The work was supported, in part, by the U.S. National Science Foundation, the U.S. National Aeronautics and Space Administration (NASA), the Swiss Federal Office for the Environment, the VoLo Foundation, the United Kingdom Natural Environment Research Council, and the Korea Meteorological Administration Research and Development Program.
MIT study reveals a new role for cell membranes Long thought to be mainly a structural support, the cell membrane also influences how cells respond to signals and may contribute to the growth of cancer cells.Cells are enveloped by a lipid membrane that gives them structure and provides a barrier between the cell and its environment. However, evidence has recently emerged suggesting that these membranes do more than simply provide protection — they also influence the behavior of the protein receptors embedded in them.
A new study from MIT chemists adds further support to that idea. The researchers found that changing the composition of the cell membrane can alter the function of a membrane receptor that promotes proliferation.
Epidermal growth factor receptor (EGFR) can be locked into an overactive state when the cell membrane has a higher than normal concentration of negatively charged lipids, the researchers found. This may help to explain why cancer cells with high levels of those lipids enter a highly proliferative state that allows them to divide uncontrollably.
“The longstanding dogma of what a membrane does is that it’s just a scaffold, an organizational structure. However, there have been increasing observations that suggest that maybe these membrane lipids are actually playing a role in receptor function,” says Gabriela Schlau-Cohen, the Robert T. Haslam and Bradley Dewey Professor of Chemistry at MIT and the senior author of the study.
The findings open up the possibility of discovering new ways to treat tumors by neutralizing the negative charge, which might turn down EGFR signaling, she adds.
Shwetha Srinivasan PhD ’22 is the lead author of the paper, which appears in the journal eLife. Other authors include former MIT postdocs Xingcheng Lin and Raju Regmi, Xuyan Chen PhD ’25, and Bin Zhang, an associate professor of chemistry at MIT.
Receptor dynamics
The EGF receptor, which is found on cells that line body surfaces and organs, is one of many receptors that help control cell growth. Some types of cancer, especially lung cancer and glioblastoma, overexpress the EGF receptor, which can lead to uncontrolled growth.
Like most receptor proteins, EGFR spans the entire cell membrane. Until recently, it has been challenging to study how signals are conveyed across the entire receptor, because of the difficulty of creating membranes that have proteins going all the way through them and then studying both ends of those proteins.
To make it easier to study these signaling processes, Schlau-Cohen’s lab uses nanodiscs, a special type of self-assembling membrane that mimics the cell membrane. When making these discs, the researchers can embed receptors in them, allowing the team to study the function of the full-length receptor.
Using a technique called single molecule FRET (fluorescence resonance energy transfer), the researchers can study how the shape of the receptor changes under different conditions. Single molecule FRET allows them to measure the distance between different parts of the protein by labeling them with fluorescent tags and then measuring how fast energy travels between the tags.
In previous work, Schlau-Cohen and Zhang used single molecule FRET and molecular dynamics simulations to reveal what happens when EGFR binds to EGF. They found that this binding causes the transmembrane section of the receptor to change shape, and that shape-shift triggers the section of the receptor that extends inside the cell to activate cellular machinery that stimulates growth.
Stuck in an overactive state
In the new study, the researchers used a similar approach to investigate how altering the composition of the membrane affects the function of the receptor. First, they explored how elevated levels of negatively charged lipids would affect the cell membrane and EGFR function.
Normally, about 15 percent of the cell membrane is made up of negatively charged lipids. The researchers found that membranes with negatively charged lipids in the range of 15 to 30 percent behaved normally, but if that level reached 60 percent, then the EGFR receptor would become locked into an active state.
In that state, the pro-growth signaling pathway is turned on all the time, even when no EGF is bound to the receptor. Many cancer cells show increased levels of these lipids, and this mechanism could help to explain why those cells are able to grow unchecked, Schlau-Cohen says.
“If the membrane has high levels of negatively charged lipids, then it’s always in that open conformation. It doesn’t matter if ligand is bound or unbound,” she says. “It’s always in the conformation that’s telling the cell to grow, not just when EGF binds.”
The researchers also used this system to explore the role of cholesterol in EGFR function. When the researchers created nanodiscs with elevated cholesterol levels, they found that the membranes became more rigid, and this rigidity suppressed EGFR signaling.
The research was funded by the National Institutes of Health and MIT’s Department of Chemistry.
Waves hit different on other planetsFrom lazy ripples to towering breakers, waves should vary widely from one planet to another, according to a new model.On a calm day, a light breeze might barely ripple the surface of a lake on Earth. But on Saturn’s largest moon Titan, a similar mild wind would kick up 10-foot-tall waves.
This otherworldly behavior is one prediction from a new wave model developed by scientists at MIT. The model is the first to capture the full dynamics of waves and what it takes to whip them up under different planetary conditions.
In a study published in the Journal of Geophysical Research: Planets, the MIT team introduces the model, which they’ve aptly coined “PlanetWaves.” They apply the model to predict how waves behave on planetary bodies that might host liquid lakes and oceans, including Titan, ancient Mars, and three planets beyond the solar system.
The model predicts that a gentle wind would be enough to stir up huge waves on Titan, where lakes are filled with light liquid hydrocarbons. In contrast, it would take hurricane-force winds to barely move the surface of a lake on the exoplanet 55-Cancri e, which is thought to be a lava world covered in hot, dense liquid rock.
“On Earth, we get accustomed to certain wave dynamics,” says study author Andrew Ashton, associate scientist at the Woods Hole Oceanographic Institution (WHOI) and faculty member of the MIT-WHOI Joint Program. “But with this model, we can see how waves behave on planets with different liquids, atmospheres, and gravity, which can kind of challenge our intuition.”
The team is particularly keen to understand how waves form on Titan. The large moon is the only other planetary body in the solar system other than the Earth that is known to currently host liquid lakes.
“Anywhere there’s a liquid surface with wind moving over it, there’s potential to make waves,” says Taylor Perron, the Cecil and Ida Green Professor of Earth, Atmospheric and Planetary Sciences at MIT. “For Titan, the tantalizing thing is that we don’t have any direct observation of what these lakes look like. So we don’t know for sure what kind of waves might exist there. Now this model gives us an idea.”
If humans were to one day to send a probe to Titan’s lakes, the team’s new model could inform the design of wave-resilient spacecraft.
“You would want to build something that can withstand the energy of the waves,” says lead author Una Schneck, a graduate student in MIT’s Department of Earth, Atmospheric and Planetary Sciences (EAPS). “So it’s important to know what kind of waves these instruments would be up against.”
The study’s co-authors include Charlene Detelich and Alexander Hayes of Cornell University and Milan Curcic of the University of Miami.
“The first puff”
When wind blows over water, it creates waves that can be strong enough to carve out coastlines and redistribute sediment brought to the coast by rivers. Through this process, waves can be a significant force in shaping a landscape over time. Schneck and her colleagues, who study landscape evolution on Earth and other planets, wondered how waves might behave on other worlds where gravity, atmospheric conditions, and liquid compositions can be very different from what is found on Earth.
“There have been attempts in the past to predict how gravity will affect waves on other planets,” Schneck says. “But they don’t quantify other factors such as the composition of the liquid that is making waves. That was the big leap with this project.”
She and her colleagues developed a full wave model that takes into account not just a planet’s gravity, but also properties of its surface liquid, such as its density, viscosity, and surface tension, or how resistant a liquid is to rippling. The team also incorporated the effect of a planet’s atmospheric pressure. With this model, they aimed to predict how a planet’s liquid surface would evolve in response to winds of a given speed.
“Imagine a completely still lake,” Ashton offers. “We’re trying to figure out the first puff that will make those first little tiny ripples, on up to a full ocean wave.”
Making waves
The team first tested their new model with wave data on Earth. They used measurements of waves that were collected by buoys across Lake Superior over 20 years. They found that the model, which took into account Earth’s gravity, the composition of liquid (water), and atmospheric conditions, was able to accurately predict what windspeeds it would take to generate waves across the lake, and how high the waves grew with a given wind strength.
The researchers then applied the model to predict how waves would behave on other planetary bodies that are known to host liquid on their surface. They looked first to Titan, where NASA’s Cassini mission previously captured radar images of lake formations, which scientists suspect are currently filled with liquid methane and ethane. The team used the new model to calculate the moon’s wave dynamics given its gravity, atmospheric pressure, and liquid composition.
They found that on Titan, it’s surprisingly easy to make waves. The relatively light liquid, combined with low gravity and atmospheric pressure, means that even a gentle wind can stir up huge waves.
“It kind of looks like tall waves moving in slow motion,” Schneck says. “If you were standing on the shore of this lake, you might feel only a soft breeze but you would see these enormous waves flowing toward you, which is not what we would expect on Earth.”
The researchers also considered wave activity on ancient Mars. The Red Planet hosts many impact basins that may have once been filled with water, before the planet’s atmosphere dissipated and the water evaporated away. One of those basins is Jezero Crater, which is currently being explored by NASA’s Perseverance rover. With the new model, the team showed that as Mars’ atmosphere gradually disappeared, reducing its pressure over time, it would have required stronger winds to make the same waves.
Beyond the solar system, the researchers applied the model to three different exoplanets. The first, LHS1140b, is a “cool super-Earth,” meaning that it is colder and larger than Earth. The planet hosts liquid water, though because it is so large, it has a stronger gravity. The model showed that the same wind on Earth would generate much smaller waves of water on the super-Earth, due to its difference in gravity.
The team also considered Kepler 1649b, a Venus-like planet, which has a gravity similar to Earth’s, with lakes of sulfuric acid, which is about twice as dense as water. Under these conditions, the researchers found that it would take strong winds to make even a ripple on the exo-Venus, compared to on Earth.
This effect is even more pronounced for the third planet, 55-Cancri e — a lava world that has both a higher gravity than Earth and a much denser, more viscous surface liquid. Scientists suspect that the planet hosts oceans of liquefied rock. In this environment, the model predicts that hurricane-force winds on Earth, of about 80 miles per hour, would generate only small waves of a few centimeters in height on the lava world.
Aside from illuminating new ways that waves can behave on other planets, Perron hopes the model will answer longstanding questions of planetary landscape formation.
“Unlike on Earth where there is often a delta where a river meets the coast, on Titan there are very few things that look like deltas, even though there are plenty of rivers and coasts. Could waves be responsible for this?” Perron wonders. “These are the kinds of mysteries that this model will help us solve.”
This work was supported, in part, by NASA and the National Science Foundation.
Geothermal energy turns red hotMIT Energy Initiative symposium maps a path to tap the planet’s heat-rich rocks for clean power at scale.Drill deep and drill differently. That’s what’s needed to exploit the nearly bottomless promise of geothermal energy in the United States and around the globe, according to participants at the 2026 Spring Symposium, titled “Next-generation geothermal energy for firm power.”
Sponsored by the MIT Energy Initiative (MITEI), the March 4 event drew 120 people, including MIT faculty and students, investors, and representatives from startups, multinational energy companies, and zero-carbon advocacy groups.
“The time feels right to pull together good policy, great corporate partners, and the research and technological innovations … to make significant advances in the widespread utilization of this incredible resource,” said Karen Knutson, the vice president for government affairs at MIT, in welcoming attendees.
Technology from the oil and gas industry helped usher in a first wave of geothermal energy. But chewing vertical holes through rocks in traditional ways can’t deliver on the full potential of this resource. And the real treasure — geologic formations radiating heat at 374 degrees Celsius and above — lies kilometers beneath Earth’s surface, far beyond the reach of most conventional drilling rigs.
Panelists explored the many innovations in accessing and circulating subsurface heat, as well as digging to unprecedented depths through extremely challenging geological conditions, discussing advanced drilling technologies, materials, and subsurface imaging.
This work is needed urgently, as demand for firm (always-on) power skyrockets in response to the electrification of industry and rise of data centers, said Pablo Dueñas‑Martínez, a MITEI research scientist. “We cannot get through this only with solar and wind; we need dense, deployable energy like geothermal.”
From “minuscule” to “almost inexhaustible” energy
In her opening remarks, Carolyn Ruppel, MITEI’s deputy director of science and technology, noted that despite decades of successful projects in places like the United States, Kenya, Iceland, Indonesia, and Turkey, geothermal still contributes only a “minuscule” share of global electricity. “The tremendous heat beneath our feet remains largely untouched,” she said.
Citing MIT’s milestone 2006 study “The Future of Geothermal Energy,” keynote speaker John McLennan, a professor at the University of Utah and co–principal investigator of the U.S. Department of Energy’s Utah FORGE enhanced geothermal systems (EGS) field laboratory, reminded attendees that the continental crust holds enough accessible heat to supply power for generations. “For practical purposes, it’s almost inexhaustible,” he said.
The question now, he said, is how to access that resource economically and responsibly.
At the Utah FORGE test site, McLennan has been part of a team investigating one method — adapting the oil and gas industry’s drilling and reservoir engineering expertise for hot, relatively impermeable rocks.
The project has drilled multiple deep wells into crystalline granitic rock, including a pair of wells that have been hydraulically stimulated and connected. In a recent circulation test, cold water was pumped down one well, flowed through fractures, and returned hot through the other.
“On a commercial basis … this hot water would be converted to electricity at the surface,” McLennan said. “This has now been demonstrated at Utah FORGE.”
The basic physics, in other words, work. The harder problems now are cost, repeatability, and scale.
Geothermal on the grid
Several panels highlighted the fact that next-generation geothermal is already beginning to deliver firm power.
At Lightning Dock, New Mexico, geothermal company Zanskar used a probabilistic modeling framework that simulated thousands of possible subsurface configurations to identify where to drill a new production well at an underperforming geothermal field. By thermal power delivered, the resulting well is now “the most-productive pumped geothermal well in the country,” said Joel Edwards, Zanskar’s co-founder and chief technology officer — powering the entire 15 megawatt (MW) Lightning Dock plant from a single well.
This data-driven approach enables the company to find and develop new resources faster and more cheaply than traditional methods, said Edwards.
José Bona, the director of next-generation geothermal at Turboden, explained how his company’s technology uses specialized turbines to circulate organic fluids that conserve heat better than water, and then convert that heat efficiently into electrical power. This closed-cycle technology can utilize low- to medium-temperature heat sources. Turboden is supplying its technology both to the Lightning Dock geothermal facility in New Mexcio and to Fervo Energy’s Cape Station in southwest Utah, an EGS project that will begin delivering 100 MW of baseload, clean electricity to the grid this year, aiming for 500 MW by 2028.
In Geretsried, Germany, Eavor has developed its own proprietary closed-loop system by creating a kind of underground radiator.
“We drilled to about 4.5 kilometers vertical depth, completed six horizontal multilateral pairs, and we delivered the first power to the grid in December,” said Christian Besoiu, the team lead of technology development at Eavor. The project will ultimately be capable of supplying 8.2 MW of electricity to the 32,000 households in the Bavarian town of Geretsried and 64 MW of thermal energy to the district in which the town lies, prioritizing heat when needed.
Beyond oil and gas technology
Early geothermal exploration typically targeted preexisting faults using vertical wells left by oil and gas drilling. Today, companies are experimenting with rock fracturing at multiple subsurface levels and creating heat reservoirs in previously untenable formations by using propping materials.
“Instead of vertical wells, we’re going to horizontal wells, we’re going to cased wells, we’re introducing proppants [solid materials that hold open hydraulically fractured rock] … we do dozens of stages with these designs,” said Koenraad Beckers, the geothermal engineering lead at ResFrac. This shale-style approach has already yielded much higher flow rates and more-reliable performance than earlier EGS.
Some current geothermal wells manage to achieve depths close to 15,000 feet using the oil and gas industry’s polycrystalline diamond compact drill bits, which can bore through hard rock like granite at more than 100 feet per hour. But these bits and the rigs that drive them are no match for conditions six or more kilometers down — and it is at those depths that the heat on hand begins to make an overwhelming economic case for geothermal.
“If we go to around 300 to 350 degrees, your power potential increases 10 times,” said Lev Ring, CEO of Sage Geosystems. “At that point, with reasonable CAPEX [capital expenditure] assumptions, levelized cost of electricity [a metric for comparing the cost of electricity across different generation technologies] is around 4 cents, and geothermal becomes cheaper than any other alternative.”
But “at 10 kilometers down … the largest land rigs in existence today cannot handle it,” Ring added. “We need alternatives — new materials, new ways to handle pressure, maybe even welding on the rig … a whole space that has not been addressed yet.”
One panel, featuring Quaise Energy, an MIT spinout with MITEI roots, spotlighted just how radically drilling might change. Co-founder Matt Houde described the company’s millimeter-wave drilling approach, which uses high-frequency electromagnetic waves derived from fusion research to vaporize rock instead of grinding it, as with conventional drilling. In a recent Texas field test, the team drilled 100 meters of hard basement rock in about a month, and is now planning kilometer-scale trials aimed at reaching superhot rock temperatures around 400 C, where each well could deliver many times the power of today’s geothermal projects.
Innovations for deep drilling
Moderating a panel on “MIT innovations for next-generation geothermal,” Andrew Inglis, the venture builder in residence with MIT Proto Ventures, whose position is sponsored by the U.S. Department of Energy GEODE program, framed the Institute’s role in getting such hard-tech ideas out of the lab and into the field. “The way MIT thinks about tech development, uniquely from other universities, can play a very singular role in geothermal commercial liftoff,” he said.
Materials researchers on that panel illustrated the point. Matěj Peč, an associate professor of geophysics in the Department of Earth, Atmospheric and Planetary Sciences, outlined work to build sensors that survive up to 900 C so that rock deformation and fracturing can be studied at supercritical conditions. Michael Short, the Class of 1941 Professor in the Department of Nuclear Science and Engineering, and C. Cem Tasan, the POSCO Associate Professor of Metallurgy in the Department of Materials Science and Engineering, respectively described coatings and alloys designed to resist corrosion, fouling, and cracking in extreme environments. In response to audience questions after their talks, Tasan made an important point, highlighting how academics need input from industry to understand the real-world problems (e.g., corrosion of pipes by geofluids) that require engineering solutions.
Other researchers are rethinking how to detect geothermal resources: Wanju Yuan, a research scientist with the Geological Survey of Canada at Natural Resources Canada, is using satellite imagery and thermal infrared sensing to screen vast regions for subtle hot spots and structures, processing thousands of images to identify promising sites in just a few months of work. “It’s a very efficient way to screen potential areas before more expensive exploration, thus reducing exploration and drilling risks,” he said.
Policy as backdrop, not center stage
Policy loomed in the background of many discussions — from bipartisan support for geothermal exploration and tax incentives to issues of regulation and permitting.
For Ruppel, that was by design.
“We wanted this meeting to showcase what’s technically possible and what’s already happening on the ground,” she said. “The policy world is starting to pay attention. Our job is to make sure that when that spotlight turns our way, next-generation geothermal is ready.”
MITEI’s Spring Symposium was followed by a gathering of geothermal entrepreneurs, investors, and energy industry experts co-hosted by MITEI and the Clean Air Task Force. “GeoTech Summit: Accelerating geothermal technology, projects, and deal flow” explored the financing challenges and opportunities of geothermal energy today.
MIT faculty, alumni receive 2025-26 American Physical Society honorsTwo faculty and six additional alumni win top APS awards and prizes; four faculty and 12 additional alumni named APS Fellows.The American Physical Society (APS) recently honored two MIT faculty members — professors Yoel Fink PhD ’00 and Mehran Kardar PhD ’83 — as well as six alumni with prizes and awards for their contributions to physics and academic leadership.
In addition, several MIT faculty members — Professor Jorn Dunkel, Professor Yen-Jie Lee PhD ’11, Associate Professor Mingda Li PhD ’15, and Associate Professor Julien Tailleur — as well as 12 additional alumni were named APS Fellows.
Yoel Fink PhD ’00, the Danae and Vasilis (1961) Salapatas Professor in the Department of Materials Science and Engineering, received the Andrei Sakharov Prize “for defending the academic freedom and human rights of scientists working in the U.S.”
The prize, named for physicist and human rights advocate Andrei Sakharov, recognizes scientists whose leadership and impact advance the principles of intellectual freedom and human dignity. Fink’s research focuses on “computing fabrics” — fibers and textiles that sense, communicate, store, and process information. By embedding functionality at the fiber level, fabrics become computing systems that can infer human activity and context while keeping the traditional qualities of garments. These textiles enable noninvasive monitoring of physiological and health conditions, with applications ranging from fetal and maternal health to human performance analytics, injury prevention in challenging environments, and defense.
Mehran Kardar PhD ’83, the Francis Friedman Professor of Physics, received the Lars Onsager Prize “for ground-breaking contributions to statistical physics, including the Kardar-Parisi-Zhang equation, Casimir forces, active matter, and aspects of biological physics.”
Kardar’s research focuses on how complex behavior emerges from simple interactions in systems both in and far from equilibrium, including stable ones like a still pond and rapidly changing ones such as growing surfaces. The Kardar-Parisi-Zhang equation, which he helped develop, provides a unifying framework for understanding how randomness and fluctuations shape evolving phenomena, from fluids and interfaces to biological and quantum systems. His work has also advanced the theoretical understanding of disordered materials, soft matter such as polymers and gels, and fluctuation-induced forces — including Casimir forces arising from quantum and thermal effects. More recently, he has applied these ideas to active matter — systems of self-driven units — and biological systems, helping reveal patterns in living and evolving systems.
Alumni receiving awards
Joel Butler PhD ’75 was presented the W.K.H. Panofsky Prize in Experimental Particle Physics “for wide-ranging scientific, technical, and strategic contributions to particle physics, particularly exceptional leadership in fixed-target quark flavor experiments at Fermilab and collider physics at the Large Hadron Collider.”
Anthony Duncan PhD ’75 is the recipient of the Abraham Pais Prize for History of Physics “for research on the history of quantum physics between 1900 and 1927 that culminated in 'Constructing Quantum Mechanics,' an exemplary work that uses primary sources masterfully and employs scaffold and arch metaphors to describe developments in the quantum revolution.”
Laura A. Lopez ’04 was presented the Edward A. Bouchet Award “for pioneering contributions to X-ray astronomy, including foundational studies of supernova remnants, compact objects, and stellar feedback in galaxies, and for transformative leadership in advancing equity and inclusion in physics through innovative mentorship programs, national advocacy, and unwavering support for students from historically marginalized communities.”
Zhiquan Sun PhD ’25 is the recipient of the J.J. and Noriko Sakurai Dissertation Award in Theoretical Particle Physics “for applying effective field theory to advance our understanding of QCD [quantum chromodynamics], including establishing a new formalism to study heavy quark fragmentation, determining how confinement affects energy correlators, and revealing an overlooked complexity of the axion solution to the strong CP [charge conjugation symmetry and parity symmetry] problem.”
Charles B. Thorn III ’68 received the Dannie Heineman Prize for Mathematical Physics for “fundamental contributions to elementary particle physics, primarily the theory of strong interactions and the development of string theory.”
Christina Wang ’19 received the Mitsuyoshi Tanaka Dissertation Award in Experimental Particle Physics “for pioneering a novel technique using CMS [Compact Muon Solenoid] muon chambers to search for weakly-coupled sub-GeV [giga-electronvolt] mass dark matter using long-lived particle searches, and for groundbreaking work in quantum sensing to enable new probes of dark matter.”
APS Fellows
Several MIT faculty were elected 2025 APS Fellows:
Jorn Dunkel, MathWorks Professor of Mathematics, is the recipient of the Division of Statistical and Nonlinear Physics Fellowship “for pioneering contributions to statistical, nonlinear, and biological physics, notably in understanding pattern formation in soft matter and biology, cell positioning in tissues, and turbulence in active media.”
Yen-Jie Lee PhD '11, professor of physics, received the Division of Nuclear Physics Fellowship “for pioneering measurements of jet quenching, medium response and heavy-quark diffusion in the quark-gluon plasma, and for using electron-positron collisions as an innovative control to understand collectivity in small collision systems.”
Mingda Li PhD '15, associate professor of nuclear science and engineering, is the recipient of the Topical Group on Data Science Fellowship “for pioneering the integration of artificial intelligence with scattering and spectroscopy, enabling breakthroughs in phonons, topological states, optical and time-resolved spectra, and data-driven discovery for quantum and energy applications.”
Julien Tailleur, associate professor of physics, is the recipient of the Division of Soft Matter Fellowship “for foundational theoretical work on motility-induced phase separation and emergent collective behavior in scalar active matter.”
The following additional MIT alumni were also honored as APS Fellows:
Andrew Cross SM ’05, PhD ’08 (EECS), Division of Quantum Information Fellowship
Kevin D. Dorfman SM '01, PhD '02 (ChemE), Division of Polymer Physics Fellowship
Geoffroy Hautier PhD '11 (DMSE), Division of Computational Physics Fellowship
Douglas J. Jerolmack PhD '06 (EAPS), Division of Statistical and Nonlinear Physics Fellowship
Brian Lantz '92, PhD '99 (Physics), Division of Gravitational Physics Fellowship
Valerio Lucarini SM '03 (EAPS), Topical Group on Physics of Climate Fellowship
Giles Novak '81 (Physics), Division of Astrophysics Fellowship
Steve Presse PhD '08 (Physics), Division of Biological Physics Fellowship
Jonathan Rothstein PhD '01 (MechE), Division of Fluid Dynamics Fellowship
Gray Rybka PhD '07 (Physics), Division of Particles and Fields Fellowship
Sarah Sheldon '08, PhD '13 (Physics, NSE), Forum on Industrial and Applied Physics Fellowship
Lian Shen ScD '01 (MechE), Division of Fluid Dynamics Fellowship
Learning with audiobooksA new study finds that audiobooks help students learn new words — especially when paired with one-on-one instruction.Millions of students nationwide use text-supplemented audiobooks, learning tools that are thought to help those who struggle with reading keep up in the classroom. A new study from scientists at MIT’s McGovern Institute for Brain Research finds that many students do benefit from the audiobooks, gaining new vocabulary through the stories they hear. But study participants learned significantly more when audiobooks were paired with explicit one-on-one instruction — and this was especially true for students who were poor readers. The group’s findings were reported on March 17 in the journal Developmental Science.
“It is an exciting moment in this ed-tech space,” says Grover Hermann Professor of Health Sciences and Technology John Gabrieli, noting a rapid expansion of online resources meant to support students and educators. “The admirable goal in all this is: Can we use technology to help kids progress, especially kids who are behind for one reason or another?” His team’s study — one of few randomized, controlled trials to evaluate educational technology — suggests a nuanced approach is needed as these tools are deployed in the classroom. “What you can get out of a software package will be great for some people, but not so great for other people,” Gabrieli says. “Different people need different levels of support.” Gabrieli is also a professor of brain and cognitive sciences and an investigator at the McGovern Institute.
Ola Ozernov-Palchik and Halie Olson, scientists in Gabrieli’s lab, launched the audiobook study in 2020, when most schools in the United States had closed to slow the spread of Covid-19. The pandemic meant the researchers would not be able to ask families to visit an MIT lab to participate in the study — but it also underscored the urgency of understanding which educational technologies are effective, and for whom.
“What we were really concerned about as the pandemic hit is that the types of gaps that we see widen through the summers — the summer slide that affects poor readers and disadvantaged children to a greater extent — would be amplified by the pandemic,” says Ozernov-Palchik. Many educational technologies purport to ameliorate these gaps. But, Ozernov-Palchik says, “fewer than 10 percent of educational technology tools have undergone any type of research. And we know that when we use unproven methods in education, the students who are most vulnerable are the ones who are left further and further behind.”
So the team designed a study that could be done remotely, involving hundreds of third- and fourth-graders around the country. They focused on evaluating the impact of audiobooks on children’s vocabularies, because vocabulary knowledge is so important for educational success. Ozernov-Palchik explains that books are important for exposing children to new words, and when children miss out on that experience because they struggle to read, they can fall further behind in school.
Audiobooks allow students to access similar content in a different way. For their study, the researchers partnered with Learning Ally, an organization that produces audiobooks synchronized with highlighted text on a computer screen, so students can follow along as they listen.
“The idea is, they’re going to learn vocabulary implicitly through accessing those linguistically rich materials,” Ozernov-Palchik says. But that idea was untested. In contrast, she says, “we know that really what works in education, especially for the most vulnerable students, is explicit instruction.”
Before beginning their study, Ozernov-Palchik and Olson trained a team of online tutors to provide that explicit instruction. The tutors — college students with no educational expertise — learned how to apply proven educational methods to support students’ learning and understanding of challenging new words they encountered in their audiobooks.
Students in the study were randomly assigned to an eight-week intervention. Some were asked to listen to Learning Ally audiobooks for about 90 minutes a week. Another group received one-on-one tutoring twice a week, in addition to listening to audiobooks. A third group, in which students participated in mindfulness practice without using audiobooks or receiving tutoring, served as a control.
A diverse group of students participated, spanning different reading abilities and socioeconomic backgrounds. The study’s remote design — with flexibly scheduled testing and tutoring sessions conducted over Zoom — helped make that possible. “I think the pandemic pushed researchers to rethink how we might use these technologies to make our research more accessible and better represent the people that we’re actually trying to learn about,” says Olson, a postdoc who was a graduate student in Gabrieli’s lab.
Testing before and after the intervention showed that overall, students in the audiobooks-only group gained vocabulary. But on their own, the books did not benefit everyone. Children who were poor readers showed no improvement from audiobooks alone, but did make significant gains in vocabulary when the audiobooks were paired with one-on-one instruction. Even good readers learned more vocabulary when they received tutoring, although the differences for this group were less dramatic.
Individualized, one-on-one instruction can be time-consuming, and may not be routinely paired with audiobooks in the classroom. But the researchers say their study shows that effective instruction can be provided remotely, and you don’t need highly trained professionals to do it.
For students from households with lower socioeconomic status, the researchers found no evidence of significant gains, even when audiobooks were paired with explicit instruction — further emphasizing that different students have different needs. “I think this carefully done study is a note of caution about who benefits from what,” Gabrieli says.
The researchers say their study highlights the value and feasibility of objectively evaluating educational technologies — and that effort will continue. At Boston University, where she is a research assistant professor, Ozernov-Palchik has launched a new initiative to evaluate artificial intelligence-based educational tools’ impacts on student learning.
Slice and diceSNIPE, a newly characterized biological defense system, directly protects bacteria by chopping up invading viral DNA.What if the Trojan horse had been pulled to pieces, revealing the ruse and fending off the invasion, just as it entered the gates of Troy?
That’s an apt description of a newly characterized bacterial defense system that chops up foreign DNA.
Bacteria and the viruses that infect them, bacteriophages — phages for short — are ceaselessly at odds, with bacteria developing methods to protect themselves against phages that are constantly striving to overcome those safeguards.
New research from the Department of Biology at MIT, recently published in Nature, describes a defense system that is integrated into the protective membrane that encapsulates bacteria. SNIPE, which stands for surface-associated nuclease inhibiting phage entry, contains a nuclease domain that cleaves genetic material, chopping the invading phage genome into harmless fragments before it can appropriate the host’s molecular machinery to make more phages.
Daniel Saxton, a postdoc in the Laub Lab and the paper’s first author, was initially drawn to studying this bacterial defense system in E. coli, in part because it is highly unusual to have a nuclease that localizes to the membrane, as most nucleases are free-floating in the cytoplasm, the gelatinous fluid that fills the space inside cells.
“The other thing that caught my attention is that this is something we call a direct defense system, meaning that when a phage infects a cell, that cell will actually survive the attack,” Saxton says. “It’s hard to fend off a phage directly in a cell and survive — but this defense system can do it.”
Light it up
For Saxton, the project came into focus during a fluorescence-based experiment in which viral genetic material would light up if it successfully penetrated the bacteria.
“SNIPE was obliterating the phage DNA so fast that we couldn’t even see a fluorescent spot,” Saxton recalls. “I don’t think I’ve ever seen such an effective defense system before — you can barrage the bacteria with hundreds of phage per cell, but SNIPE is like god-tier protection.”
When the nuclease domain of SNIPE was mutated so it couldn’t chop up DNA, fluorescent spots appeared as usual, and the bacteria succumbed to the phage infection.
Bacteria maintain tight control over all their defense systems, lest they be turned against their host. Some systems remain dormant until they flare up, for example, to halt all translation of all proteins in the cell, while others can distinguish between bacterial DNA and foreign, invading phage DNA. There were only two previously characterized mechanisms in the latter category before researchers uncovered SNIPE.
“Right now, the phage field is at a really interesting spot where people are discovering phage defense systems at a breakneck pace,” Saxton says.
Problems at the periphery
Saxton says they had to approach the work in a somewhat roundabout way because there are currently no published structures depicting all the steps of phage genome injection. Studying processes at the membrane is challenging: Membranes are dense and chaotic, and phage genome injection is a highly transient process, lasting only a few minutes.
SNIPE seems to discern viral DNA by interacting with proteins the phage uses to tunnel through the bacteria’s protective membrane. This “subcellular localization,” according to Saxton, may also prevent SNIPE from inadvertently chopping up the bacteria’s own genetic material.
The model outlined in the paper is that one region of SNIPE binds to a bacterial membrane protein called ManYZ, while another region likely binds to the tape measure protein from the phage.
The tape measure protein got its name because it determines the length of the phage tail — the part of the phage between the small, leglike protrusions and the bulbous head, which contains the phage’s genetic material. The researchers revealed that the phage’s tape measure protein enters the cytoplasm during injection, a phenomenon that had not been physically demonstrated before.
There may also be other proteins or interactions involved.
“If you shunt the phage genome injection through an alternate pathway that isn’t ManYZ, suddenly SNIPE doesn’t defend against the phage nearly as well,” Saxton says. “It’s unclear exactly how these proteins interact, but we do know that these two proteins are involved in this genome injection process.”
Future directions
Saxton hopes that future work will expand our understanding of what occurs during phage genome injection and uncover the structures of the proteins involved, especially the tunnel complex in the membrane through which phages insert their genome.
Members of the Laub Lab are already collaborating with another lab to determine the structure of SNIPE. In the meantime, Saxton has been working on a new defense system in which molecular mimicry — bacterial proteins imitating phage proteins — may play a role.
Michael T. Laub, the Salvador E. Luria Professor of Biology and a Howard Hughes Medical Institute investigator, notes that one of the breakthrough experiments for demonstrating how SNIPE works came from a brainstorming session at a lab retreat.
“Daniel and I were kind of stuck with how to directly measure the effect of SNIPE during infection, but another postdoc in the lab, Ian Roney, who is a co-author on the paper, came up with a very clever idea that ultimately worked perfectly,” Laub recalls. “It’s a great example of how powerful internal collaborations can be in pushing our science forward.”
Physicists zero in on the mass of the fundamental W boson particleThe team’s ultra-precise measurement confirms the Standard Model’s predictions.When fundamental particles are heavier or lighter than expected, physicists’ understanding of the universe can tip into the unknown. A particle that is just beyond its predicted mass can unravel scientists’ assumptions about the forces that make up all of matter and space. But now, a new precision measurement has reset the balance and confirmed scientists’ theories, at least for one of the universe’s core building blocks.
In a paper appearing today in the journal Nature, an international team including MIT physicists reports a new, ultraprecise measurement of the mass of the W boson.
The W boson is one of two elementary particles that embody the weak force, which is one of the four fundamental forces of nature. The weak force enables certain particles to change identities, such as from protons to neutrons and vice versa. This morphing is what drives radioactive decay, as well as nuclear fusion, which powers the sun.
Now, scientists have determined the mass of the W boson by analyzing more than 1 billion proton-colliding events produced by the Large Hadron Collider (LHC) at CERN (the European Organization for Nuclear Research) in Switzerland. The LHC accelerates protons toward each other at close to the speed of light. When they collide, two protons can produce a W boson, among a shower of other particles.
Catching a W boson is nearly impossible, as it decays almost immediately into two types of particles, one of which, a neutrino, is so elusive that it cannot be detected. Scientists are left to measure the other particle, known as a muon, and model how it might add up to the total mass of its parent, the W boson. In the new study, scientists used the Compact Muon Solenoid (CMS) experiment, a particle detector at the LHC that precisely tracks muons and other particles produced in the aftermath of proton collisions.
From billions of proton-proton collisions, the team identified 100 million events that produced a W boson decaying to a muon and a neutrino. For each of these events, they carried out detailed analyses to narrow in on a precise mass measurement. In the end, they determined that the W boson has a mass of 80360.2 ± 9.9 megaelectron volts (MeV). This new mass is in line with predictions of the Standard Model, which is physicists’ best rulebook for describing the fundamental particles and forces of nature.
The precision of the new measurement is on par with a previous measurement made in 2022 by the Collider Detector at Fermilab (CDF). That measurement took physicists by surprise, as it was significantly heavier than what the Standard Model predicted, and therefore raised the possibility of “new physics,” such as particles and forces that have yet to be discovered.
Because the new CMS measurement is just as precise as the CDF result and agrees with the Standard Model along with a number of other experiments, it is more likely that physicists are on solid ground in terms of how they understand the W boson.
“It’s just a huge relief, to be honest,” says Kenneth Long, a lead author of the study, who is a senior postdoc in MIT’s Laboratory for Nuclear Science. “This new measurement is a strong confirmation that we can trust the Standard Model.”
The study is authored by more than 3,000 members of CERN’s CMS Collaboration. The core group who worked on the new measurement includes about 30 scientists from 10 institutions, led by a team at MIT that includes Long; Tianyu Justin Yang PhD ’24; David Walter and Jan Eysermans, who are both MIT postdocs in physics; Guillelmo Gomez-Ceballos, a principal research scientist in the Particle Physics Collaboration; Josh Bendavid, a former research scientist; and Christoph Paus, a professor of physics at MIT and principal investigator with the Particle Physics Collaboration.
Piecing together
The W boson was first discovered in 1983 and is predicted to be the fourth heaviest among all the fundamental particles. Multiple experiments have aimed to narrow in on the particle’s mass, with varying degrees of precision. For the most part, these experiments have produced measurements that agree with the Standard Model’s predictions. The 2022 measurement by Fermilab’s CDF experiment is the one significant outlier. It also happens to be the most precise experiment to date.
“If you take the CDF measurement at face value, you would say there must be physics beyond the Standard Model,” says co-author Christoph Paus. “And of course that was the big mystery.”
Paus and his colleagues sought to either support or refute the CDF’s findings by making an independent measurement, with an experiment that matches CDF’s precision. Their new W boson mass measurement is a product of 10 years’ worth of work, both to analyze actual particle collision events and to simulate all the scenarios that could produce those events.
For their new study, the physicists analyzed proton collision events that were produced at the LHC in 2016. When it is running, the particle collider generates proton collisions at a furious rate of about one every 25 nanoseconds. The team analyzed a portion of the LHC’s 2016 dataset that encompasses billions of proton-proton collisions. Among these, they identified about 100 million events that produced a very short-lived W boson.
“A particle like the W boson exists for a teeny tiny moment — something like 10-24 seconds — before decaying to two particles, one of which is a neutrino that can’t be measured directly,” Long explains. “That’s the tricky part: You have to measure the other particle — a muon — really well, and be able to piece things together with only one piece of the puzzle.”
Gathering momentum
When a muon is produced from the decay of a W boson, it carries half of the W boson’s mass, which is converted into momentum that carries the muon away from the original collision. Due to the strong magnetic field inside the CMS detector, the electrically charged muon follows a path whose curvature is a function of its momentum. Scientists’ challenge is to track the muon’s path and every interaction it may have with other particles and its surroundings, in order to estimate its initial momentum.
The muon’s momentum is also influenced by the momentum of the W boson before it decays. Decoding the impact of the W boson’s motion from the effects of its mass presented a major challenge. To infer the W boson mass, the team first carried out simulations of every scenario they could think of that a muon might experience after a proton-proton collision in the chaotic environment of the particle collider. In all, the team produced 4 billion such simulated events described by state-of-the-art theoretical calculations. The simulations encoded diverse hypotheses about how the muon momentum is affected by the physical features of the CMS detector, as well as uncertainties in the predictions that govern W boson production in LHC collisions.
The researchers compared their simulations with data from the 2016 LHC run. For every proton-proton collision event that occurs in the collider, scientists can use the CMS detector at CERN’s LHC to precisely measure the energy and momentum of resulting particles such as muons. The team analyzed CMS measurements of muons that were produced from over 100 million W boson events. They then overlaid this data onto their simulations of the muon momentum, which they then converted to a new mass for the W boson.
That mass — 80360.2 ± 9.9 megaelectron volts — is significantly lighter than the CDF experiment’s measurement. What’s more, the new estimate is within the range of what the Standard Model predicts for the W boson’s mass, bolstering physicists’ confidence in the Standard Model and its descriptions of the major particles and forces of nature.
“With the combination of our really precise result and other experiments that line up with the Standard Model’s predictions, I think that most people would place their bets on the Standard Model,” Long says. “Though I do think people should continue doing this measurement. We are not done.”
“We want to add more data, make our analysis techniques more precise, and basically squeeze the lemon a little harder. There is always some juice left,” Paus adds. “With a better look, then we can say for certain whether we truly understand this one fundamental building block.”
This work was supported, in part, by multiple funding agencies, including the U.S. Department of Energy, and the SubMIT computing facility, sponsored by the MIT Department of Physics.
Two physicists and a curious host walk into a studio…On GBH’s new show The Curiosity Desk, MIT LIGO researchers revel in the beauties of fundamental discovery science and MIT astronomers talk planetary defense.This March on The Curiosity Desk, GBH’s daily science show with host Edgar B. Herwick III, MIT scientists dropped by to address the questions: “How close are we to observing the dark universe?” (Thursday, March 12 episode) and “Is Earth prepared for asteroids?” (Thursday, March 26 episode).
Up first, Prof. Nergis Mavalvala, dean of the MIT School of Science, and Prof. Salvatore Vitale joined the host live in studio to talk about the science behind the Laser Interferometer Gravitational-wave Observatory (LIGO) and how LIGO has provided the ability to observe the universe in ways that have never been done before.
In addition to learning something new, Mavalvala explained how experimenting delivers an added piece of excitement: “pushing the technology, the precision of the instrument, requires you to be very inventive. There’s almost nothing in these experiments that you can go buy off a shelf. Everything you’re designing, everything is from scratch. You’re meeting very stringent requirements.”
Herwick likened how they might tweak or tinker with the experiment to souping up a car engine, and the LIGO scientists nodded – adding that in the most complex experiments, each bite-sized part on its own works well, and it’s the interfaces between them that scientists must get right.
While there, the two long-time colleagues also took a detour to explain how in physics experimentalists benefit from the work of theorists and vice versa. Mavalvala, whose work focuses on building the world’s most precise instruments to study physical phenomena, described the synergy between ideas that come from theory (work that Vitale does) and how you measure. (No, they assure Herwick, they don’t get into a lot of fights.)
In fact, it’s fantastic to have people from both worlds at MIT, said Vitale. Mavalvala agreed. “One of the things that’s really important about theory in science is that ultimately, in physics especially, it’s a bunch of math. And the important thing that you have to ask is, ‘does nature really behave that way?’ And how do you answer that question? You have to go out and measure. You have to go observe nature,” said Mavalvala.
As scientists fine-tune the gravitational wave detectors, they will inform what data are collected, what astrophysical objects they might find or hope to find – and the search for certain fainter, farther away, or more exotic objects can inform what enhancements they prioritize.
But what if I’m not interested in any of that, asked Herwick? Why should I care?
“To me, it falls in the category of for the betterment of humankind. You never know what is going to be useful. A lot of fundamental research was very far at the beginning from what turned out to be fundamental applications,” said Vitale, adding, “What they do on the instrument side has already now very important applications.”
Mavalvala was unequivocal, underscoring how pursuing curiosity is put to good use:
“When you’re making instruments that achieve that kind of precision, you’re inventing new technologies. [With LIGO] We’ve invented vibration isolation technologies to keep our mirrors really still. We’ve invented lasers that are quieter than any that were ever made before. We’ve invented photonic techniques that are allowing us to make applications even to far off things like quantum computing.
“So, this is one of the beauties of fundamental discovery science. A, you’ll discover something. But B you’ll be doing two things: you’ll be inventing the technologies of the future, and you’ll be training the generations of scientists who may go off to do completely different things, but this is what inspires them.”
Watch the full conversation below and on YouTube:
Planetary defense
Turning to objects beyond Earth – specifically, asteroids – Associate Professor Julien de Wit, along with research scientists Artem Burdanov and Saverio Cambioni, joined Herwick at the Curiosity Desk later in the month. They talked about their ongoing research to identify smaller asteroids (about the size of a school bus) using the James Webb Space Telescope and why planetary defense goes beyond thinking about the massive asteroids featured in movies like Armageddon. Notably, a lot of technology on earth depends on satellites, and asteroids pose the biggest threat to satellites.
“Dinosaurs didn’t need to care about an asteroid hitting the moon. Humanity a century ago didn’t care. Now, if [an asteroid] hits the moon, a lot of debris will be expelled and all those particles – big and small – they will affect the fleet of satellites around Earth. That’s a big potential problem, so we need to take that into account in our future,” said Burdanov.
There’s also a potential upside to being better able to detect and potentially “capture” asteroids, explained de Wit, all of it benefitted by new instruments. “It’s really an asteroid revolution going on… Our situational awareness of what’s out there is really about to change dramatically.”
He explains that one dream is to mine asteroids themselves for material to build or power next generation technologies or stations in space. “The way to reliably move into space is to use resources from space. We can’t just move stuff to build a full city. We use stuff from space.”
Echoing the sentiments expressed earlier in the month by MIT’s dean of science, the trio of asteroid explorers also described how the pursuits of planetary scientists can lead to unexpected rewards along the way. “We are swimming in an era that is data rich, and so what we do in our group and at MIT is mine that data to reveal the universe like never before,” says de Wit. “Revealing new populations of asteroids, new populations of planets, and making sense of our universe like we have never done.”
Watch the full conversation below and on the GBH YouTube channel:
Tune in to the Curiosity Desk some Thursdays to hear from MIT researchers as they visit Herwick and the production team.
Climate change may produce “fast-food” phytoplanktonWith warmer ocean temperatures, the composition of marine plankton could shift from protein-rich to carb-heavy, a new study suggests.We are what we eat. And in the ocean, most life-forms source their food from phytoplankton. These microscopic, plant-like algae are the primary food source for krill, sea snails, some small fish, and jellyfish, which in turn feed larger marine animals that are prey for the ocean’s top predators, including humans.
Now MIT scientists are finding that phytoplankton's composition, and the basic diet of the ocean, will shift significantly with climate change.
In an open-access study appearing today in the journal Nature Climate Change, the team reports that as sea surface temperatures rise over the next century, phytoplankton in polar regions will adapt to be less rich in proteins, heavier in carbohydrates, and lower in nutrients overall.
The conclusions are based on results from the team’s new model, which simulates the composition of phytoplankton in response to changes in ocean temperature, circulation, and sea ice coverage. In a scenario in which humans continue to emit greenhouse gases through the year 2100, the team found that changing ocean conditions, particularly in the polar regions, will shift phytoplankton’s balance of proteins to carbohydrates and lipids by approximately 20 percent. The researchers analyzed observations from the past several decades, and already have found a signature of this change in the real world.
“We’re moving in the poles toward a sort of fast-food ocean,” says lead author and MIT postdoc Shlomit Sharoni. “Based on this prediction, the nutritional composition of the surface ocean will look very different by the end of the century.”
The study’s MIT co-authors are Mick Follows, Stephanie Dutkiewicz, and Oliver Jahn; along with Keisuke Inomura of the University of Rhode Island; Zoe Finkel, Andrew Irwin, and Mohammad Amirian of Dalhousie University in Halifax, Canada; and Erwan Monier of the University of California at Davis.
Nutritional information
Phytoplankton drift through the upper, sun-lit layers of the ocean. Like plants on land, the marine microalgae are photosynthetic. Their growth depends on light from the sun, carbon dioxide from the atmosphere, and nutrients such as nitrogen and iron that well up from the deep ocean.
When studying how phytoplankton will respond to climate change, scientists have primarily focused on how rising ocean temperatures will affect phytoplankton populations. Whether and how the plankton’s composition will change is less well-understood.
“There’s been an awareness that the nutritional value of phytoplankton can shift with climate change,” says Sharoni, “But there has been very little work in directly addressing that question.”
She and her colleagues set out to understand how ocean conditions influence phytoplankton macromolecular composition. Macromolecules are large molecules that are essential for life. The main types of macromolecules include proteins, lipids, carbohydrates, and nucleic acids (the building blocks of DNA and RNA). Every form of life, including phytoplankton, is composed of a balance of macromolecules that helps it to survive in its particular environment.
“Nearly all the material in a living organism is in these broad molecular forms, each having a particular physiological function, depending on the circumstances that the organism finds itself in,” says Follows, a professor in the Department of Earth, Atmospheric and Planetary Sciences.
An unbalanced diet
In their new study, the researchers first looked at how today’s ocean conditions influence phytoplankton’s macromolecular composition. The team used data from lab experiments carried out by their collaborators at Dalhousie. These experiments revealed ways in which phytoplankton’s balance of macromolecules, such as proteins to carbohydrates, shifted in response to changes in water temperature and the availability of light and nutrients.
With these lab-based data, the group developed a quantitative model that simulates how plankton in the lab would readjust its balance of proteins to carbohydrates under different light and nutrient conditions. Sharoni and Inomura then paired this new model with an established model of ocean circulation and dynamics developed previously at MIT. With this modeling combination, they simulated how phytoplankton composition shifts in response to ocean conditions in different parts of the world and under different climate scenarios.
The team first modeled today’s current climate conditions. Consistent with observations, their model predicts that that a little more than half of the average phytoplankton cell today is composed of proteins. The rest is a mix of carbohydrates and lipids.
Interestingly, in polar regions, phytoplankton are slightly more protein-rich. At the poles, the cover of sea ice limits the amount of sunlight phytoplankton can absorb. The researchers surmise that phytoplankton may have adapted by making more light-harvesting proteins to help the organisms efficiently absorb the weak sunlight.
However, when they modeled a future climate change scenario, the team found a significant shift in phytoplankton composition. They simulated a scenario in which humans continue to emit greenhouse gases through the year 2100. In this scenario, the ocean sea surface temperatures will rise by 3 degrees Celsius, substantially reducing sea ice coverage. Warmer temperatures will also limit the ocean’s circulation, as well as the amount of nutrients that can circulate up from the deep ocean.
Under these conditions, the model predicts that the population of phytoplankton growth in polar regions will increase significantly, consistent with earlier studies. Uniquely, this model predicts that phytoplankton in polar regions will shift from a protein-rich to a carb- and lipid-heavy composition. They found that plankton will not need as much light-harvesting protein, since less sea ice will make sunlight more easily available for the organisms to absorb. Total protein levels in these polar phytoplankton will decline by up to 30 percent, with a corresponding increase in the contribution of carbs and lipids.
It’s unclear what impact a larger population of carb- and lipid-heavy phytoplankton may have on the rest of the marine food web. While some organisms may be stressed by a reduction in protein, others that make lipid stores to survive through the winter might thrive.
The team also simulated phytoplankton in subtropical, higher-latitude regions. In these ocean areas, it’s expected that phytoplankton populations will decline by 50 percent. And the team’s modeling shows that their composition will also shift.
With warmer temperatures, the ocean’s circulation will slow down, limiting the amount of nutrients that can upwell from the deep ocean. In response, subtropical phytoplankton may have to find ways to live at deeper depths, to strike a balance between getting enough sunlight and nutrients. Under these conditions, the organisms will likely shift to a slightly more protein-rich composition, making use of the same photosynthetic proteins that their polar counterparts will require less of.
On balance, given the projected changes in phytoplankton populations with climate change, their average composition around the world will shift to a more carb-heavy, low-nutrient composition.
The researchers went a step further and found that their modeling agrees with available small set of actual phytoplankton field samples that other scientists previously collected from Arctic and Antarctic regions. These samples showed compositions of phytoplankton have become more carb- and lipid-heavy over the past few decades, as the team’s model predicts under climate warming.
“In these regions, you can already see climate change, because sea ice is already melting,” Sharoni explains. “And our model shows that proteins in polar plankton have been declining, while carbs and lipids are increasing.”
“It turns out that climate change is accelerated in the Arctic, and we have data showing that the composition of phytoplankton has already responded,” Follows adds. “The main message is: The caloric content at the base of the marine food web is already changing. And it’s not a clear story as to how this change will transmit through the food web.”
This work was supported, in part, by the Simons Foundation.
QS World University Rankings rates MIT No. 1 in 12 subjects for 2026The Institute also ranks second in seven subject areas.QS World University Rankings has placed MIT in the No. 1 spot in 12 subject areas for 2026, the organization announced today.
The Institute received a No. 1 ranking in the following QS subject areas: Chemical Engineering; Chemistry; Civil and Structural Engineering; Computer Science and Information Systems; Data Science and Artificial Intelligence; Electrical and Electronic Engineering; Engineering and Technology; Linguistics; Materials Science; Mechanical, Aeronautical, and Manufacturing Engineering; Mathematics; and Physics and Astronomy.
MIT also placed second in seven subject areas: Architecture/Built Environment; History of Art; Biological Sciences; Economics and Econometrics; Marketing; Natural Sciences; and Statistics and Operational Research.
For 2026, universities were evaluated in 55 specific subjects and five broader subject areas.
Quacquarelli Symonds Limited subject rankings, published annually, are designed to help prospective students find the leading schools in their field of interest. Rankings are based on research quality and accomplishments, academic reputation, and graduate employment.
MIT has been ranked as the No. 1 university in the world by QS World University Rankings for 14 straight years.
Alex Tang’s dream of becoming a physician started in grade school when he read Lisa Sanders’ “Diagnosis” column in The New York Times Magazine. Although he often encountered unfamiliar medical terms, Tang was captivated by the magic of medicine, as Sanders described how physicians turned puzzling sets of symptoms into concrete diagnoses and treatment plans for patients.
A decade later, Tang is one step closer to achieving his dream. The MIT senior has challenged himself academically, dual-majoring in chemistry and biology and minoring in biomedical engineering. “All of the courses have encouraged me to think about problems through different lenses,” he says.
Tang has also challenged himself as the editor-in-chief of MIT’s student newspaper, The Tech, and as a competitive triathlete. In the fall, he will begin medical school, where he hopes to develop clinical skills and continue honing his scientific abilities. Ultimately, he aspires to pursue a career as a physician-scientist, focusing on how cancers respond to and resist treatment. He wants to help convert those insights into novel therapies that can be tailored to individual cancer patients.
“I want to advance precision oncology, ensuring that each patient receives the most effective, personalized treatment possible,” he says.
Thriving in the lab
Originally from Massachusetts, Tang was eager to make the most of his MIT experience, especially because of its extensive research opportunities. “Both my parents worked in the Cambridge biotech space, and being able to contribute to innovative science here has been a priority,” he says.
Early on, Tang gravitated toward oncology after joining the Nir Hacohen Lab at the Broad Institute, an interest cemented after taking 7.45 (Cancer Biology), which was taught by professors Tyler Jacks and Michael Hemann. Fascinated by how new cancer therapies were changing patients’ lives, he joined a project with implications for patients with difficult prognoses: For the last three-and-half years, Tang has been studying the effects of combined immunotherapy and targeted molecular therapy on tumors in patients with metastatic colorectal cancer.
“I hope my work can provide clarity for patients and physicians, and empower them to be confident in their options for care,” Tang says.
Last year, Tang was awarded a prestigious Goldwater Scholarship, which supports undergraduates who go on to become leading scientists, engineers, and mathematicians in their respective fields.
In addition to gaining technical skills, Tang has found working in the Hacohen Lab to be enriching in other important ways.
“What’s been great about research is learning from experts in the field who become your role models,” he says, “They are at the frontiers of investigating the most challenging questions in the field, and iterating through the scientific process with them is such a joy.”
Looking forward to medical school, he hopes to complement his basic science research with work that is more clinically involved.
“I want to bridge the gap between fundamental discoveries and tangible improvements in patient care,” Tang says. He has already set out on this mission, recently leading the development of a prognostic assay in lung cancer.
Breaking news
After stopping by the booth for MIT’s student newspaper, The Tech, during Campus Preview Weekend, Tang knew he wanted to join and contribute to a publication that has long chronicled MIT’s history and culture. Starting as a news writer and later serving as editor-in-chief, he learned how to write under pressure, reported on major campus events, and balanced leadership with collaboration.
“It’s been such an honor and pleasure to document people across the diverse MIT community who are all contributing to the character of the Institute in different ways,” he says.
It’s an activity he’ll drop everything for.
“When we have things come up and we have to do a breaking news story or we have some editorial thing that needs to be managed, I’ll just stop working to sort out whatever’s happening,” he says. “I think that’s what passion really is about.”
His journey with The Tech has not always been easy. In the summer between his first and second year, he found himself solely responsible for producing the paper’s news content amidst a staff shortage while the paper was facing financial difficulties.
“Coming into sophomore fall, I focused on recruiting more staff and seeking out ways to get more funding,” Tang says. “The paper wouldn’t be here without the people, both students and faculty advisors alike, who bought into The Tech’s mission.”
Though he hopes to pursue a career in medicine, Tang has found journalism to be integral in shaping how he will connect and communicate with patients and colleagues.
“You are responsible for taking someone’s story, breaking it down, and retelling it in your own words in a way that you feel would resonate with the audience and serve the community,” he says.
An outlet through triathlon
Despite his busy schedule, Tang prioritizes staying active and maintaining fitness. A former competitive swimmer in high school and now a triathlete, he still finds himself drawn back to the water when everything around him feels fast-paced.
“Swimming, biking, and running are good ways to de-stress,” Tang says. “It’s therapeutic in the sense that you can just let go. The race is just that culmination of letting it go at a more elevated level.”
He credits MIT’s infrastructure for helping him stay committed to training. “My dorm is steps away from the pool and the track,” he says. “The convenience is superb.”
Tang has found success in competitions, most recently placing third in his age group at the 2025 Boston Triathlon. In fact, it is the feeling of accomplishment that pushes him every day.
“There are many days when you want to take it easy, but you have to remember the joy waiting for you at the end of the race when you’ve put in the work,” he says. “It motivates me to be conscious and aware of what I’m doing in practice.”
During the summer, Tang and his younger brother go out for long runs in the Boston suburbs. “It is great to have my brother push me every day,” Tang says. “There has been no one more supportive of me than my family.”
A complicated future for a methane-cleansing moleculeA new model shows how levels of the “atmosphere’s detergent” may rise and fall in response to climate change.Methane is a powerful greenhouse gas that is second only to carbon dioxide in driving up global temperatures. But it doesn’t linger in the atmosphere for long thanks to molecules called hydroxyl radicals, which are known as the “atmosphere’s detergent” for their ability to break down methane. As the planet warms, however, it’s unclear how the air-cleaning agents will respond.
MIT scientists are now shedding some light on this. The team has developed a new model to study different processes that control how levels of hydroxyl radical will shift with warming temperatures.
They find that the picture is complicated. As temperatures increase, so too will water vapor in the atmosphere, which will in turn boost the molecule’s concentrations. But rising temperatures will also increase “biogenic volatile organic compound emissions” — gases that are naturally released by some plants and trees. These natural emissions can reduce hydroxyl radical and dampen water vapor’s boosting effect.
Specifically, the team finds that if the planet’s average temperatures rise by 2 degrees Celsius, the accompanying rise in water vapor will increase hydroxyl radical levels by about 9 percent. But the corresponding increase in biogenic emissions would in turn bring down hydroxyl radical levels by 6 percent. The final accounting could mean a small boost, of about 3 percent, in the atmosphere’s ability to break down methane and other chemical compounds as the planet warms.
“Hydroxyl radicals are important in determining the lifetime of methane and other reactive greenhouse gases, as well as gases that affect public health, including ozone and certain other air pollutants,” says study author Qindan Zhu, who led the work as a postdoc in MIT’s Department of Earth, Atmospheric and Planetary Sciences (EAPS).
“There’s a whole range of environmental reasons why we want to understand what’s going on with this molecule,” adds Arlene Fiore, the Peter H. Stone and Paola Malanotte Stone Professor in EAPS. “We want to make sure it’s around to chemically remove all these gases and pollutants.”
Fiore and Zhu’s new study appears today in the Journal of Advances in Modeling Earth Systems (JAMES). The study’s MIT co-authors include Jian Guan and Paolo Giani, along with Robert Pincus, Nicole Neumann, George Milly, and Clare Singer of Lamont-Doherty Earth Observatory and the Columbia Climate School, and Brian Medeiros at the National Center for Atmospheric Research.
A natural neutralizer
The hydroxyl radical, known chemically as OH, is made up of one oxygen atom and one hydrogen atom, along with an unpaired electron. This configuration makes the molecule extremely reactive. Like a chemical vacuum cleaner, OH easily pulls an electron or hydrogen atom away from other molecules, breaking them down into weaker, more water-soluble forms. In this way, OH reduces a vast range of chemicals, including some air pollutants, pathogens, and ozone. And changes in OH are a powerful lever on methane.
“For methane, the reaction with OH is considered the most important loss pathway,” Zhu says. “About 90 percent of the methane that’s removed from the atmosphere is due to the reaction with OH.”
Indeed, it’s thanks to reactions with hydroxyl radical that methane can only stick around in the atmosphere for about a decade — far shorter than carbon dioxide, which can linger for 1,000 years or longer. But even as OH breaks down methane already in the atmosphere, more methane continues to accumulate. Rising methane concentrations, in addition to human-derived emissions of carbon dioxide, are driving global warming, and it’s unclear how OH’s methane-clearing power will keep up.
“The questions we’re exploring here are: What are the main processes that control OH concentrations? And how will OH respond to climate change?” Fiore says.
An aquaplanet’s air
For their study, the researchers developed a new model to simulate levels of OH in the atmosphere under a current global climate scenario, compared to a future warmer climate. Their model, dubbed “AquaChem,” is an expansion of a simplified model that is part of a suite of tools developed by the Community Earth System Model (CESM) project. The model that the team chose to build off is one that represents the Earth as a simplified “aquaplanet,” with an entirely ocean-covered surface.
Aquaplanet models allow scientists to study detailed interactions in the atmosphere in response to changes in surface temperatures, without having to also spend computing time and energy on simulating complex dynamics between the land, water, and polar ice caps.
To the aquaplanet model, Zhu added an atmospheric chemistry component that simulates detailed chemical reactions in the atmosphere consistent with the applied surface temperatures. The chemical reactions that she modeled represent those that are known to affect OH concentrations.
OH is primarily produced when ozone interacts with sunlight in the presence of water vapor. For instance, scientists have found that OH levels can vary depending certain anthropogenic and natural emissions, all of which Zhu incorporated separately and together into the AquaChem model in order to isolate the impact of each process on OH.
The emissions in particular include carbon monoxide, methane, nitrogen oxides, and volatile organic compounds (VOCs), some of which are emitted through human practices, and others that are given off by natural processes. One type of naturally-derived VOCs are “biogenic” emissions — gases, such as isoprene, that some plants and trees emit through tiny pores called stomata during transpiration.
Into the AquaChem model, Zhu plugged in data that were available for each type of emissions from the year 2000 — a year that is generally considered to represent the current climate in a simplified form. She set the aquaplanet’s sea surface temperatures to the zonal annual mean of that year, and found that the model accurately reproduced the major sensitivities of OH chemistry to the underlying chemical processing as simulated in a more complex chemistry-climate model.
Then, Zhu ran the model under a second, globally warming scenario. She set the planet’s sea surface temperatures to warm by 2 degrees Celsius (a warming that is likely to occur unless global anthropogenic carbon emissions are mitigated). The team looked at how this warming would affect the various types of emissions and chemical processes, and how these changes would ultimately affect levels of OH in the atmosphere.
In the end, they found the two biggest drivers of OH levels were rising water vapor and biogenic emissions. They found that global warming would increase the amount of water vapor to the atmosphere, which in turn would boost production of OH by 9 percent. However, this same degree of warming would also increase biogenic emissions such as isoprene, which reacts with and breaks down OH, bringing down its levels by 6 percent.
The team recognizes that there are many other factors that affect the response of isoprene emissions to surface warming. Rising CO2, not considered in this study, may dampen this temperature-driven response. Of all the factors that can shift OH levels under global warming, the researchers caution that biogenic emissions are the most uncertain, even though they appear to have a large influence. Going forward, the scientists plan to update AquaChem to continue studying how biogenic emissions, as well as other processes and climate scenarios, could sway OH concentrations.
“We know that changes in atmospheric OH, even of a few percent, can actually matter for interpreting how methane might accumulate in the atmosphere,” Zhu says. “Understanding future trends of OH will allow us to determine future trends of methane.”
This work was supported, in part, by Spark Climate Solutions and the National Oceanic and Atmospheric Administration.